Fast-Scan Cyclic Voltammetry for Adenosine Detection: Principles, Advances, and Clinical Translation
This article provides a comprehensive resource for researchers and drug development professionals on the application of Fast-Scan Cyclic Voltammetry (FSCV) for detecting the neuromodulator adenosine.
Fast-Scan Cyclic Voltammetry for Adenosine Detection: Principles, Advances, and Clinical Translation
Abstract
This article provides a comprehensive resource for researchers and drug development professionals on the application of Fast-Scan Cyclic Voltammetry (FSCV) for detecting the neuromodulator adenosine. It covers foundational principles, from adenosine's role as a signaling molecule to its electrochemical signature. The review details methodological innovations in waveform design, electrode engineering, and data analysis that enhance sensitivity and selectivity. It further addresses key troubleshooting challenges such as electrode fouling and ohmic drop, and evaluates FSCV against other neurochemical techniques. Finally, it explores the validation pathway and growing potential for clinical application of FSCV in understanding neurological disorders and developing closed-loop therapeutic systems.
Adenosine as a Neuromodulator and Its Electrochemical Fundamentals
The Biological Role of Adenosine in Sleep, Blood Flow, and Neurotransmission
Adenosine is a purine nucleoside that functions as a ubiquitous signaling molecule and inhibitory neurotransmitter throughout the body [1] [2]. It serves as a fundamental cellular component with diverse regulatory functions in physiological processes including sleep-wake regulation, cerebral blood flow, and neuromodulation [3] [4] [5]. As a breakdown product of adenosine triphosphate (ATP), adenosine provides a critical link between cellular energy metabolism and neuronal activity [3] [6]. Recent advances in detection methodologies, particularly fast-scan cyclic voltammetry (FSCV), have revealed novel rapid signaling modes of adenosine action in the brain, opening new avenues for investigating its complex roles in health and disease [7] [4] [8]. This application note details the biological significance of adenosine and provides standardized protocols for its detection in experimental settings, with particular emphasis on FSCV methodologies relevant to current neuroscience research.
Biological Functions and Mechanisms of Action
Adenosine in Sleep-Wake Regulation
Adenosine functions as an endogenous sleep-promoting substance that accumulates in the brain during wakefulness and gradually declines during sleep [3] [5]. The sleep-promoting effects of adenosine are mediated through multiple receptor mechanisms and brain regions:
- Receptor Mechanisms: Adenosine promotes sleep primarily through A1 and A2A receptors. A1 receptor activation inhibits wake-promoting neurons in the basal forebrain, while A2A receptor stimulation in the ventrolateral preoptic area promotes sleep [5].
- Cellular Energy Link: As neuronal activity consumes ATP during wakefulness, extracellular adenosine levels rise as a byproduct of ATP metabolism, creating a homeostatic sleep drive that correlates with prior wakefulness duration [3] [6].
- Caffeine Interaction: The stimulant effects of caffeine occur primarily through non-selective antagonism of adenosine receptors, particularly A1 and A2A subtypes, thereby blocking adenosine's sleep-inducing effects [3] [5] [1].
Table 1: Adenosine Concentrations Under Different Physiological Conditions
| Condition | Brain Region | Adenosine Concentration | Biological Significance |
|---|---|---|---|
| Normal extracellular levels | Multiple | 165-300 nM [6] | Basal neuromodulation |
| Normal plasma levels | Blood | 0.04-0.2 μM [2] | Circulating adenosine pool |
| Cellular damage/ischemia | Inflammatory or ischemic tissue | 600-1200 nM [1] | Cytoprotective signaling |
| Prolonged wakefulness | Basal forebrain | Significantly elevated [5] | Sleep homeostasis |
Adenosine in Cardiovascular Regulation
Adenosine exerts potent effects on the cardiovascular system, with both physiological regulatory and clinical applications:
- Coronary Vasodilation: Adenosine causes dilation of coronary arteries, making it valuable for myocardial perfusion imaging in patients unable to undergo exercise stress testing [1].
- Heart Rate Regulation: It decreases heart rate and can induce transient atrioventricular (AV) node block, enabling its use in terminating supraventricular tachycardias [1].
- Receptor-Mediated Effects: These cardiovascular effects are primarily mediated through A1 receptors in the AV node and A2A receptors on vascular smooth muscle [1].
Adenosine as a Neuromodulator
In the central nervous system, adenosine functions as a ubiquitous neuromodulator with predominantly inhibitory effects on neuronal activity:
- Glutamate and Dopamine Regulation: Transient adenosine release inhibits both glutamate and dopamine release via A1 receptor activation, with effects spatially constrained within approximately 250 μm in the caudate-putamen [7].
- Presynaptic Inhibition: Adenosine acts through presynaptic A1 receptors to limit adenyl cyclase formation and promote neuronal hyperpolarization, reducing neurotransmitter release probability [7] [5].
- Activity-Dependent Release: Adenosine release can be evoked by electrical or mechanical stimulation, or occur spontaneously without stimulation, suggesting multiple release mechanisms [4].
Figure 1: Adenosine Signaling Pathways and Physiological Effects. This diagram illustrates the primary mechanisms through which neuronal activity leads to adenosine accumulation and subsequent activation of receptor-mediated physiological responses including sleep promotion, neurotransmission inhibition, and vasodilation.
Experimental Detection of Adenosine Using FSCV
Principles of Fast-Scan Cyclic Voltammetry
Fast-scan cyclic voltammetry has emerged as a powerful technique for detecting rapid adenosine dynamics in the brain with sub-second temporal resolution [4] [8]. Key advantages of FSCV for adenosine detection include:
- High Temporal Resolution: Capable of detecting transient adenosine release events lasting only a few seconds [4].
- Chemical Selectivity: Provides unique cyclic voltammograms for adenosine, enabling discrimination from other electroactive compounds [8].
- Real-time Monitoring: Allows observation of spontaneous and evoked adenosine transients in living tissue [4] [8].
Table 2: FSCV Parameters for Adenosine Detection
| Parameter | Recommended Setting | Alternative Parameters | Purpose |
|---|---|---|---|
| Scan Rate | 400 V/s [8] | - | Optimal electron transfer kinetics |
| Waveform | -0.4 V to +1.45 V [8] | -0.4 V to +1.3 V (catecholamines) [9] | Covers adenosine oxidation potentials |
| Repetition Rate | 10 Hz [8] | - | Balances temporal resolution and stability |
| Holding Potential | -0.4 V [8] | - | Maintains electrode sensitivity |
| Primary Oxidation Peak | ~1.3 V [8] | - | Characteristic adenosine signature |
| Secondary Oxidation Peak | Grows over time [8] | - | Confirms adenosine detection |
Advanced FSCV Methodologies
Recent technological advances have enhanced FSCV capabilities for adenosine research:
- Multiplexed Detection: Simultaneous measurement of adenosine with other neurotransmitters (dopamine, glutamate) by combining FSCV with genetically encoded fluorescent sensors [7].
- Structural Similarity Index (SSIM) Analysis: Image-based analysis of FSCV color plots that achieves 99.5% precision and 95% recall in detecting spontaneous adenosine events [8].
- Cone-Shaped Carbon Fiber Microelectrodes: 30 μm cone-shaped electrodes demonstrate improved longevity (4.7-fold increase) and reduced glial activation compared to conventional 7 μm electrodes [9].
Research Reagent Solutions
Table 3: Essential Reagents and Materials for Adenosine Research
| Reagent/Material | Specifications | Experimental Function |
|---|---|---|
| Carbon Fiber Microelectrodes (CFMEs) | 7-30 μm diameter, 100 μm exposed length [9] [8] | Working electrode for FSCV measurements |
| Adenosine Reference Standard | High-purity (>95%), prepared in 0.1 M HClO4 [8] | Analytical standard for calibration and validation |
| A1 Receptor Antagonist (DPCPX) | 8-cyclopentyl-1,3-dipropylxanthine, selective A1 antagonist [7] | Pharmacological validation of adenosine mechanisms |
| Artificial Cerebrospinal Fluid (aCSF) | pH 7.4, containing 1.2 mM MgCl₂, 1.2 mM CaCl₂ [8] | Physiological buffer for in vitro and in vivo studies |
| Sindbis Viral Vectors | Expressing iGluSnFR3.v857 [7] | Enables expression of genetically encoded sensors for multiplexing |
| Enzyme Inhibitors | Dipyridamole (adenosine deaminase inhibitor) [1] [2] | Increases endogenous adenosine levels |
Standardized Protocol: Multiplexed Detection of Adenosine, Dopamine, and Glutamate
Figure 2: Experimental Workflow for Multiplexed Neurotransmitter Detection. This protocol outlines the sequential steps for simultaneous measurement of adenosine, dopamine, and glutamate release in brain slices using combined FSCV and fluorescence imaging techniques.
Detailed Methodology
Objective: To simultaneously characterize adenosine modulation of dopamine and glutamate release in caudate-putamen brain slices.
Materials Preparation:
- Prepare acute brain slices (300-400 μm thickness) from rodent caudate-putamen region.
- Express the genetically encoded glutamate sensor iGluSnFR3.v857 using Sindbis viral vectors 18-24 hours prior to experimentation [7].
- Fabricate carbon-fiber microelectrodes (CFMEs) from T-650 carbon fibers (7-μm diameter) with 100 μm exposed length, insulated in glass capillaries [8].
Instrumentation Setup:
- Configure FSCV system with a ChemClamp potentiostat using a two-electrode system (CFME working electrode and Ag/AgCl reference electrode).
- Set FSCV waveform parameters: -0.4 V holding potential, +1.45 V switching potential, 400 V/s scan rate, 10 Hz repetition rate [8].
- Establish fluorescence imaging system compatible with iGluSnFR3.v857 excitation/emission characteristics.
Experimental Procedure:
- Implant CFME in the caudate-putamen region near cells expressing iGluSnFR3.v857.
- Apply electrical stimulation (biphasic pulses, 300 μA, 2 ms per phase) to evoke neurotransmitter release.
- Apply exogenous adenosine locally to the brain slice for 30 seconds while monitoring neurotransmitter dynamics.
- For receptor mechanism studies, pre-perfuse with A1 receptor antagonist DPCPX (8-cyclopentyl-1,3-dipropylxanthine) before adenosine application.
- Record simultaneously using FSCV (adenosine and dopamine) and fluorescence imaging (glutamate) for 10 minutes post-adenosine application to monitor recovery.
Data Analysis:
- Process FSCV data using Structural Similarity Index (SSIM) analysis to identify adenosine events with high precision and recall [8].
- Quantify spatial propagation of adenosine effects by analyzing inhibition radius.
- Calculate recovery kinetics of dopamine and glutamate release following adenosine application.
Expected Outcomes:
- Transient inhibition of both electrically stimulated dopamine and glutamate release by adenosine.
- Spatial restriction of adenosine inhibition within approximately 250 μm radius.
- Complete blockade of adenosine effects by A1 receptor antagonist DPCPX.
- Recovery of dopamine and glutamate release within 10 minutes after adenosine application [7].
Adenosine serves as a crucial biological regulator at the intersection of sleep homeostasis, vascular function, and neuromodulation. The development of sophisticated detection methodologies, particularly FSCV and multiplexed sensing approaches, has revealed complex spatial and temporal dynamics of adenosine signaling in the brain. The standardized protocols presented herein provide researchers with robust methodologies for investigating adenosine mechanisms in physiological and pathological conditions. Continued refinement of these techniques, including improved electrode designs and advanced data analysis algorithms, will further enhance our understanding of adenosine's diverse biological roles and therapeutic potential.
Uncovering Rapid Adenosine Signaling with FSCV
Adenosine is a critical signaling molecule and neuromodulator in the central nervous system, regulating numerous physiological processes including neurotransmission, blood flow, sleep, and neuroprotection [4] [10]. Traditional understanding positioned adenosine as a slow-acting modulator operating on time scales of minutes to hours. However, recent advances in electrochemical detection methods have revealed a novel mode of rapid adenosine signaling that occurs within seconds, suggesting adenosine may function similarly to classical neurotransmitters like dopamine in certain contexts [4] [11].
Fast-scan cyclic voltammetry (FSCV) has emerged as a powerful analytical technique for detecting these rapid adenosine fluctuations with sub-second temporal resolution. This capability has transformed our understanding of adenosine dynamics, enabling researchers to characterize spontaneous, transient adenosine release in addition to stimulated release patterns [11]. The application of FSCV to adenosine detection provides unprecedented insights into the rapid modulatory roles of this purine signaling molecule, its interaction with other neurotransmitter systems, and its potential implications for neurological disorders and therapeutic development.
Electrochemical Fundamentals of Adenosine Detection
Principles of FSCV for Adenosine
FSCV detects electroactive neurotransmitters by applying a triangular waveform between a carbon-fiber microelectrode and a reference electrode. For adenosine detection, the standard waveform scans from -0.4 V to +1.45 V or +1.5 V and back versus a Ag/AgCl reference electrode at a rate of 400 V/s [10] [11]. This scan takes less than 10 ms and is repeated every 100 ms, providing a temporal resolution of 10 measurements per second. The fast scan rates generate a substantial background charging current due to double-layer charging at the electrode interface, but this stable background can be subtracted to reveal the faradaic current from analyte oxidation and reduction [10].
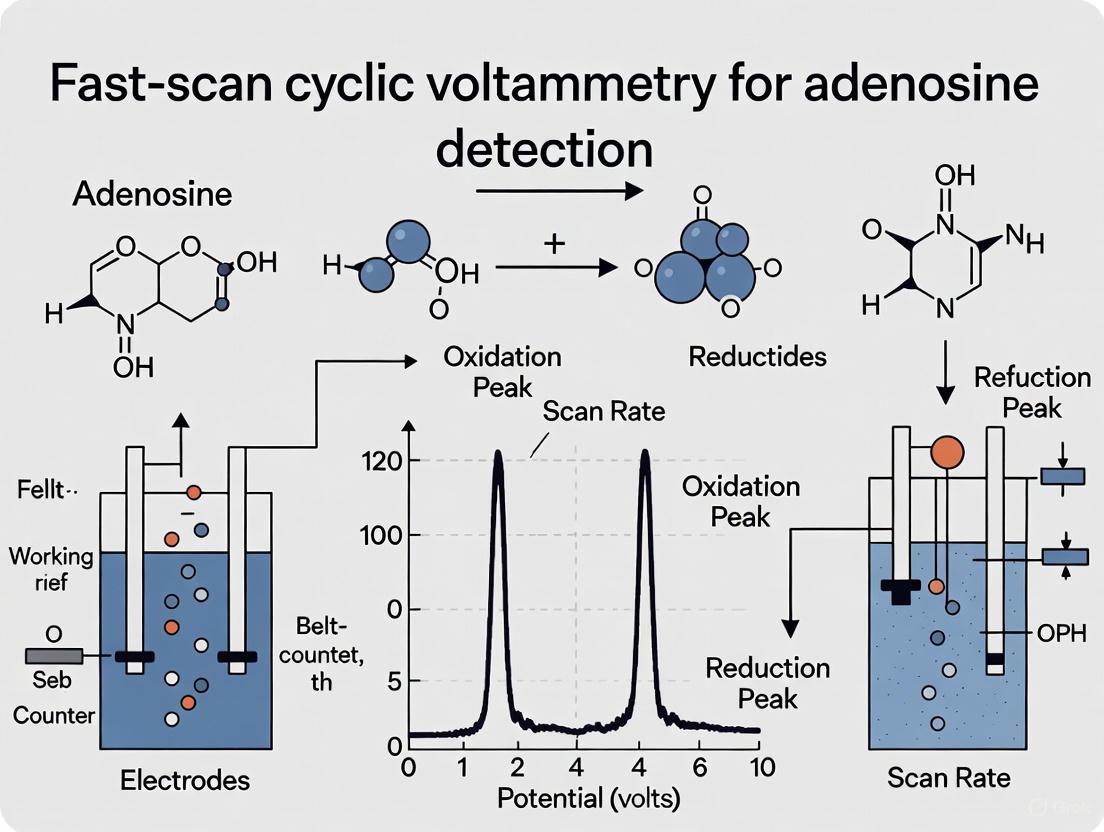
Adenosine undergoes a series of electrochemical oxidations that generate its characteristic cyclic voltammogram. The initial oxidation occurs at approximately 1.4 V, with a secondary oxidation detected at 1.0 V [10]. These first two oxidation steps are irreversible, and reduction peaks are not typically observed under standard FSCV conditions. The characteristic cyclic voltammogram for adenosine therefore displays two oxidation peaks, with the most prominent peak appearing near the switching potential at 1.4 V [10].
Data Visualization and Analysis
FSCV data are commonly visualized using false color plots, which display multiple voltammograms collected over time in a two-dimensional format [10]. A vertical slice through the color plot at any time point yields the cyclic voltammogram for that moment, while a horizontal slice shows how the oxidation current at a specific potential changes over time. The primary oxidation peak for adenosine appears approximately half a second before the secondary peak on color plots, providing a distinctive temporal signature that helps distinguish adenosine from other electroactive compounds [10].
Table 1: Key Electrochemical Parameters for Adenosine Detection with FSCV
| Parameter | Typical Setting | Notes |
|---|---|---|
| Waveform | Triangular | -0.4 V to +1.45/+1.5 V |
| Scan Rate | 400 V/s | Standard for adenosine |
| Scan Frequency | 10 Hz | 100 ms intervals |
| Primary Oxidation Peak | ~1.4 V | Irreversible |
| Secondary Oxidation Peak | ~1.0 V | Irreversible |
| Working Electrode | Carbon fiber microelectrode | 7 μm diameter typical |
| Reference Electrode | Ag/AgCl | Implanted in contralateral hemisphere |
Experimental Protocols for Adenosine Detection
Carbon-Fiber Microelectrode Preparation
CFMEs are fabricated according to established protocols [12] [11]. A single carbon fiber (typically 7 μm diameter for conventional electrodes) is aspirated into a glass capillary tube, which is then heated and pulled to form a sealed microelectrode. The protruding carbon fiber is trimmed to a length of approximately 65-100 μm using a scalpel under microscopic guidance. Electrical connection is established by applying silver paint to a lead wire inserted into the glass tube. Before experimental use, electrodes are preconditioned using FSCV sweeps to stabilize their electrochemical response [12] [11].
Recent advances in electrode design have demonstrated that larger diameter carbon fibers (30 μm) with cone-shaped modifications can significantly improve mechanical durability and signal longevity. These cone-shaped electrodes are created using electrochemical etching where a direct current voltage (typically 10 V) is applied to a carbon fiber segment submerged in Tris buffer, with a linear actuator gradually withdrawing the electrode to form the tapered conical shape [12].
In Vivo FSCV Recordings
For in vivo adenosine measurements, animals are typically anesthetized and placed in a stereotaxic frame. A burr hole is drilled at the appropriate coordinates for the brain region of interest (e.g., motor cortex or dorsal striatum) [11]. The carbon-fiber microelectrode is positioned in the target region, while an Ag/AgCl reference electrode is implanted in the contralateral hemisphere. The electrodes are connected to a voltammetric amplifier, and recordings are made by continuously applying the triangular waveform every 100 ms [11].
Following electrode implantation, an equilibration period of at least 30 minutes is recommended before initiating recordings to allow the electrochemical background current to stabilize. Adenosine transients can be recorded spontaneously or in response to various stimuli. For pharmacological validation, receptor antagonists or enzyme inhibitors can be administered systemically or locally to verify the identity of the measured signals [11].
Table 2: Characteristic Adenosine Transients Detected In Vivo
| Brain Region | Spike Amplitude | Spike Frequency | Inter-Spike Interval |
|---|---|---|---|
| Motor Cortex | 85 ± 11 nM | 0.5 - 1.5 Hz | 1 - 5 seconds |
| Dorsal Striatum | 66 ± 7 nM | 0.5 - 1.5 Hz | 1 - 5 seconds |
Signal Validation and Identification
Proper identification of adenosine signals is crucial for data integrity. Adenosine transients are identified by their characteristic cyclic voltammogram with oxidation peaks at 1.4 V and 1.0 V [10] [11]. The temporal pattern of these peaks in false color plots provides additional confirmation. Pharmacological validation using adenosine receptor antagonists or adenosine-degrading enzymes can further confirm signal identity, as these manipulations should alter the detected transients in predictable ways [10].
Several molecules with similar electrochemical properties may potentially interfere with adenosine detection. ATP contains the same electroactive adenine moiety as adenosine and generates similar oxidation signals. However, standard carbon-fiber microelectrodes with a -0.4 V holding potential show greater sensitivity for adenosine than ATP [10]. Hydrogen peroxide oxidizes at approximately 1.2 V but lacks the secondary oxidation peak characteristic of adenosine. The use of modified waveforms, such as the "sawhorse" waveform, can further enhance selectivity for adenosine over potential interferents [10].
Essential Research Tools and Reagents
Table 3: Research Reagent Solutions for Adenosine FSCV
| Item | Function/Role | Specifications |
|---|---|---|
| Carbon Fiber Microelectrodes | Working electrode for FSCV measurements | 7 μm standard; 30 μm for improved durability; cone-shaped for reduced tissue damage |
| Tris Buffer | Electrochemical stability for in vitro calibration | pH 7.4; 15 mM Trizma phosphate, 3.25 mM KCl, 140 mM NaCl, 1.2 mM CaCl2, 1.25 mM NaH2PO4, 1.2 mM MgCl2, 2.0 mM Na2SO4 |
| Adenosine Stock Solution | Analytic for calibration and experiments | 10 mM in 0.1 M perchloric acid; diluted to working concentrations in Tris buffer |
| Ag/AgCl Reference Electrode | Stable reference potential | Placed in contralateral hemisphere during in vivo recordings |
| Electrochemical Etching System | Fabrication of cone-shaped electrodes | 10 V DC in Tris buffer; linear actuator for controlled withdrawal |
| FSCV Data Acquisition System | Waveform application and current measurement | Commercial systems (e.g., WaveNeuro) or custom setups with National Instruments hardware and LabVIEW software |
Experimental Workflows and Signaling Pathways
The following diagrams illustrate the key experimental workflows and signaling concepts for adenosine detection using FSCV.
FSCV Adenosine Detection Workflow
Adenosine Signaling Pathway
Electrode Optimization Strategy
Comparison with Alternative Methodologies
FSCV offers distinct advantages and limitations compared to other techniques for adenosine measurement. Microdialysis coupled with HPLC provides excellent chemical specificity and the ability to measure multiple analytes simultaneously, but its temporal resolution is limited to minutes rather than seconds [10]. While microdialysis can detect basal adenosine levels and slow changes during events like ischemia, it cannot capture the rapid transients that FSCV reveals [10].
Enzyme-based biosensors offer good specificity for adenosine with a response time of approximately 2 seconds, bridging the gap between microdialysis and FSCV [10]. These biosensors utilize enzyme cascades that metabolize adenosine to hydrogen peroxide, which is detected amperometrically. However, FSCV remains the fastest available method for direct adenosine detection, with a sampling rate of 10 Hz enabling sub-second resolution of adenosine dynamics [10].
Electrophysiological recordings provide indirect information about adenosine signaling by monitoring its effects on neuronal firing, with millisecond temporal resolution [10]. These measurements reveal functional consequences of adenosine release but do not directly quantify adenosine concentrations. The combination of FSCV with electrophysiology at the same electrode represents a powerful approach for correlating adenosine dynamics with neuronal activity [10].
Applications and Future Directions
The application of FSCV to adenosine detection has revealed novel aspects of purine signaling in both health and disease. Research has demonstrated spontaneous, transient adenosine release in brain regions including the motor cortex, dorsal striatum, and spinal cord [11]. These transients occur with remarkable regularity at intervals of 1-5 seconds and frequencies of 0.5-1.5 Hz, suggesting they may represent a fundamental mode of adenosine signaling previously undetected by slower measurement techniques [11].
FSCV studies have also characterized activity-dependent adenosine release evoked by electrical stimulation, mechanical stimulation, and external stimuli such as tail pinch [11]. This rapid adenosine signaling appears to modulate other neurotransmitter systems, including dopamine release, and may regulate local oxygen availability [4] [10]. These findings suggest adenosine serves both rapid neuromodulatory functions in addition to its established role as a slow homeostatic regulator.
Future applications of adenosine FSCV include investigating its role in neurological disorders such as Parkinson's disease, epilepsy, and sleep disorders, where adenosine signaling is implicated but not fully understood [11]. The potential integration of FSCV with deep brain stimulation (DBS) systems represents another promising direction, enabling real-time monitoring of adenosine release during therapeutic stimulation [12] [13]. Technical advances in electrode materials, waveform design, and data analysis methods will continue to enhance the sensitivity, selectivity, and temporal resolution of FSCV for adenosine detection, further expanding our understanding of this versatile signaling molecule.
The Unique Electrochemical Fingerprint of Adenosine on Carbon Surfaces
Within the field of fast-scan cyclic voltammetry (FSCV) for neurotransmitter detection, the identification of individual analytes, particularly within complex mixtures, remains a significant challenge. While much of the data analysis focus has historically been on dopamine, there is a growing need for robust methods to detect other crucial neuromodulators. This application note details the unique electrochemical fingerprint of adenosine on carbon surfaces, providing researchers and drug development professionals with detailed protocols for its detection and analysis. Adenosine is a purine nucleoside that functions as a potent neuromodulator, regulating physiological processes such as cerebral blood flow, sleep-wake cycles, and neurotransmission. Its detection is complicated by its multi-step oxidation process and the spontaneous, unpredictable nature of its release in vivo. Unlike neurotransmitters with a single, stable cyclic voltammogram (CV) shape, adenosine exhibits a dynamic CV evolution, where its secondary oxidation peak intensifies over time, presenting a unique identification challenge that requires analysis of the entire three-dimensional data structure. This protocol focuses on leveraging the structural similarity (SSIM) index for image-based analysis of FSCV color plots, a method that significantly improves the accuracy and automation of adenosine detection.
The Electrochemical Signature of Adenosine
Adenosine produces a distinct electrochemical profile on carbon-fiber microelectrodes (CFMEs) that differentiates it from other electroactive species in the brain.
Characteristic Voltammetric Peaks
When subjected to a standard FSCV waveform (−0.4 V to +1.45 V, 400 V/s, 10 Hz), adenosine exhibits two primary oxidation peaks. The primary oxidation peak occurs at approximately 1.3 V, which is a consistent feature across adenosine events. A secondary oxidation peak emerges over time, growing in intensity as the detection event progresses. This temporal evolution of the CV shape is a hallmark of adenosine's electrochemical behavior and a critical differentiator from potential interferents [8].
Selectivity Against Common Interferents
The uniqueness of the adenosine fingerprint allows it to be effectively distinguished from other biological compounds. The SSIM-based analysis method has demonstrated excellent selectivity by successfully rejecting signals from:
- pH changes, a common source of false positives in FSCV
- Histamine, a high-oxidation potential compound
- Hydrogen peroxide (H₂O₂), another molecule with overlapping oxidation potentials [8]
Table 1: Key Characteristics of the Adenosine Electrochemical Fingerprint
| Feature | Description | Significance for Detection |
|---|---|---|
| Primary Oxidation Peak | ~1.3 V vs. Ag/AgCl | Consistent, primary identifying feature |
| Secondary Oxidation Peak | Grows in over time | Confirms identity and differentiates from static CVs |
| Color Plot Pattern | Unique 3D structure in FSCV plots | Enables image-based analysis (SSIM) |
| Dynamic CV Evolution | Voltammogram shape changes during a single release event | Distinguishes from molecules like dopamine |
Quantitative Detection Protocol
This section provides a detailed methodology for detecting adenosine using Structural Similarity Index (SSIM) analysis of FSCV color plots.
Materials and Equipment
Research Reagent Solutions
Table 2: Essential Reagents and Materials
| Item | Specification/Composition | Function/Purpose |
|---|---|---|
| Carbon-Fiber Microelectrodes (CFMEs) | T-650 carbon fibers, 7-μm diameter, 100 μm exposed length [8] | Working electrode for FSCV measurements |
| Phosphate-Buffered Saline (PBS) | 131.25 mM NaCl, 3.00 mM KCl, 10.0 mM NaH₂PO₄, 1.2 mM MgCl₂, 2.0 mM Na₂SO₄, 1.2 mM CaCl₂, pH 7.4 [8] | Physiological buffer for in vitro and in vivo studies |
| Adenosine Stock Solution | 10 mM in 0.1 M HClO₄ [8] | Stable stock for preparing working standards |
| Adenosine Triphosphate (ATP) | 10 mM in 0.1 M HClO₄ [8] | For interaction and selectivity studies |
| Potentiostat | ChemClamp with two-electrode system (CFME working electrode and Ag/AgCl reference) [8] | Instrumentation for applying waveform and measuring current |
Instrumental Parameters
The following parameters are critical for optimizing adenosine detection:
- Waveform: Triangular waveform with a holding potential of −0.4 V and a switching potential of +1.45 V
- Scan Rate: 400 V/s
- Repetition Rate: 10 Hz
- Data Collection: Use HDCV Analysis software or equivalent [8]
SSIM-Based Analysis Workflow
The following diagram illustrates the complete workflow for adenosine detection using the SSIM method, from data acquisition to final identification:
Data Preprocessing
- Apply Digital Filtering: Process the raw FSCV color plot using a second-order Butterworth high-pass filter with a half-power frequency of 0.03 Hz. This step removes background charging current and low-frequency signal drift, eliminating the need for traditional background subtraction [8].
- Smoothing (Optional): For particularly noisy data, apply a Savitzky-Golay filter (window length of 15) to smooth the data without significantly distorting the signal [8].
SSIM Calculation and Analysis
- Reference Selection: Utilize a library of validated adenosine transient references. The "Standard Library" version of the software includes 15 pre-loaded adenosine references, while the "Internal Reference" version requires the user to identify the start time of 6 transient adenosine events from their own data [8].
- Image Normalization: Normalize both the sample transient and reference color plots to their respective maximum currents before SSIM calculation. This ensures comparison is based on pattern recognition rather than absolute current intensity [8].
- SSIM Index Computation: The SSIM index between a sample image (x) and reference image (y) is calculated as a function of three components:
- Luminance (l): Comparison of mean intensity
- Contrast (c): Comparison of standard deviation of intensity
- Structure (s): Comparison of standardized intensity The overall index is given by: SSIM(x,y) = [l(x,y)] · [c(x,y)] · [s(x,y)], producing a value between 0 (no similarity) and 1 (identical) [8].
- Event Identification: Apply an optimized SSIM cutoff score to distinguish adenosine events from noise and other chemicals. Events scoring above the threshold are classified as adenosine release.
Performance Metrics and Validation
The SSIM method for adenosine detection has been rigorously validated, demonstrating high accuracy and reliability:
Table 3: Performance Metrics of SSIM-Based Adenosine Detection
| Metric | Performance Value | Interpretation |
|---|---|---|
| Precision | 99.5 ± 0.6% | Extremely high confidence that detected events are truly adenosine |
| Recall | 95 ± 3% | Captures the vast majority of genuine adenosine events |
| F1 Score | 97 ± 2% | Excellent overall balance between precision and recall |
| Data Source | 15 experiments from 3 researchers | Demonstrates method robustness across users |
Advanced Applications and Simultaneous Detection
The SSIM image analysis approach is a generalizable strategy that extends beyond adenosine detection alone.
Multi-Analyte Detection
The same SSIM framework can be optimized to detect dopamine by using dopamine-specific reference color plots. This enables the identification of simultaneous adenosine and dopamine release events, providing unprecedented insight into neuromodulator interactions in the brain [8].
Machine Learning Integration
For complex mixtures with highly similar electrochemical profiles, such as adenosine phosphates (AMP, ADP, ATP), machine learning analysis of spectral data (e.g., from Surface-Enhanced Raman Scattering) can provide an additional layer of discrimination. While not part of the core FSCV protocol, this represents a complementary approach for challenging analytical scenarios [14].
Troubleshooting and Technical Notes
- Low SSIM Scores: Ensure the high-pass filter is properly applied to remove background drift, which can obscure the adenosine signal.
- High False Positive Rate: Re-optimize the SSIM cutoff threshold for your specific experimental setup. Verify that the reference library matches the experimental conditions.
- Poor Signal-to-Noise Ratio: Confirm electrode condition and implement Savitzky-Golay smoothing as needed.
- Inconsistent Peak Potentials: Regularly calibrate the Ag/AgCl reference electrode to maintain potential stability.
This protocol establishes SSIM-based image analysis as a powerful, automated method for detecting adenosine using its unique electrochemical fingerprint on carbon surfaces. By leveraging the full three-dimensional FSCV color plot data, researchers can achieve highly accurate, efficient identification of adenosine dynamics in complex biological environments.
Adenosine is a purinergic neuromodulator that regulates critical physiological processes, including neurotransmission, blood flow, and sleep, on time scales from minutes to seconds [4]. The detection of rapid, transient adenosine release is essential for understanding its role in neuroimmune signaling and pathologies such as stroke and Parkinson's disease [15] [16]. Fast-scan cyclic voltammetry (FSCV) has emerged as a premier technique for monitoring adenosine with sub-second temporal resolution in vivo [4] [15]. However, achieving reliable analytical performance requires overcoming three interconnected challenges: sensitivity (detecting low nanomolar concentrations), selectivity (distinguishing adenosine from electroactive interferents), and fouling (maintaining sensor performance against surface contamination) [16] [8] [17]. This Application Note details the core methodologies and advanced solutions enabling robust adenosine detection within a research context focused on FSCV.
Key Challenges and Quantitative Comparisons
The table below summarizes the primary challenges in adenosine detection and the efficacy of corresponding advanced solutions, providing a performance overview for researchers.
Table 1: Key Challenges and Advanced Solutions in Adenosine Detection with FSCV
| Challenge | Description & Impact | Advanced Solutions | Demonstrated Efficacy |
|---|---|---|---|
| Sensitivity | Low basal concentrations (nanomolar range); standard carbon fiber microelectrodes (CFMEs) exhibit low sensitivity for high-oxidation-potential analytes like adenosine [16]. | Carbon Nanospikes (CNSs): Enhance surface roughness and oxide concentration [17].PEDOT:Nafion Composites: Improve charge transfer and signal uniformity [16]. | CNSs increased normalized sensitivity for adenosine by 4.8-fold vs. traditional CFMEs [17]. |
| Selectivity | Similar oxidation potentials of adenosine, H₂O₂, and histamine lead to overlapping signals [8]. | Structural Similarity Index (SSIM) Analysis: An image-based algorithm that analyzes the entire FSCV color plot [8].CNS Electrodes: Promote unique secondary products for adenosine and histamine, altering their electrochemical "fingerprints" [17]. | SSIM achieved 99.5% precision and 95% recall for identifying adenosine transients, effectively rejecting pH changes and interferents [8]. |
| Fouling | Adsorption of proteins or polymerization of oxidation products blocks the electrode surface, degrading signal over time [16] [17]. | CNS Coatings: Elevated edge planes and hydrophilic surface prevent adsorption of fouling agents [17].Plasma Treatment: Increases surface hydrophilicity using oxygen or nitrogen plasma [16]. | CNS electrodes demonstrated significantly reduced fouling for histamine and when used in brain tissue [17]. |
The Scientist's Toolkit: Essential Research Reagents and Materials
The following table catalogs key materials critical for developing and executing successful FSCV experiments for adenosine detection.
Table 2: Essential Research Reagents and Materials for Adenosine FSCV
| Item | Function/Application | Key Characteristics |
|---|---|---|
| Carbon Fiber Microelectrodes (CFMEs) | Traditional working electrode for FSCV; typically 7µm diameter [8] [18]. | Biocompatible; minimal tissue damage; low background currents; PAN-based fibers (e.g., T-650) offer fast electron transfer [18]. |
| Carbon Nanospikes (CNSs) | Nanostructured electrode coating to enhance sensitivity and impart antifouling properties [17]. | High surface roughness; abundant edge planes and oxygen functional groups; hydrophilic surface [17]. |
| PEDOT:Nafion Composite | Conductive polymer coating to improve signal stability, selectivity for cations, and biofouling resistance [16]. | Uniform coating; enhanced charge transfer; repels anionic interferents; stable in vivo for up to 6 hours [16]. |
| Tris Buffer or Artificial Cerebrospinal Fluid (aCSF) | Electrochemical cell electrolyte and calibration medium [12] [8]. | Tris buffer ensures electrochemical stability during calibration. aCSF more closely mimics the ionic composition of in vivo environments [12]. |
| Exonuclease I (exo I) | Enzyme used in signal amplification strategies for specific biosensors (e.g., ATP detection) [19]. | Cleaves single-stranded DNA (ssDNA) from the 3' terminus; enables target recycling and signal amplification [19]. |
| SSIM Analysis Software | Automated, high-accuracy data analysis tool for identifying adenosine transients from FSCV color plots [8]. | Uses image recognition on 3D FSCV data; compatible with MATLAB; high precision and recall for adenosine [8]. |
Experimental Protocols
Protocol: Fabrication and Testing of Carbon Nanospike Microelectrodes (CNSMEs) for Enhanced Adenosine Sensitivity
This protocol outlines the creation of CNS-modified electrodes, which provide superior sensitivity and antifouling properties for adenosine detection [17].
1. Electrode Fabrication: - Base Electrode Preparation: Fabricate a standard carbon fiber microelectrode (CFME) by aspirating a single carbon fiber (e.g., T-650, 7 µm diameter) into a glass capillary. Pull the capillary using a micropipette puller to form a tight seal and trim the exposed fiber to a length of ~100 µm [18]. - CNS Synthesis: Use a chemical vapor deposition (CVD) system to grow carbon nanospikes directly onto the exposed carbon fiber. A typical process involves using a hydrocarbon precursor (e.g., C₂H₂) in a hydrogen/argon atmosphere at elevated temperatures (e.g., 700-800°C) to catalyze the vertical growth of sharp, graphitic nanostructures.
2. Electrochemical Characterization and Calibration: - Setup: Use a standard two-electrode system with the CNSME as the working electrode and an Ag/AgCl wire as the reference electrode. Perform all experiments in a flow injection analysis apparatus connected to a syringe pump. - FSCV Parameters: Apply a triangular waveform with a holding potential of -0.4 V, a switching potential of 1.45 V, a scan rate of 400 V/s, and a repetition rate of 10 Hz [8]. - Calibration: Prepare a 1 mM stock solution of adenosine in 0.1 M HClO₄. Dilute to desired concentrations (e.g., 1, 2, 5 µM) in phosphate-buffered saline (PBS, pH 7.4). Inject aliquots into a continuous flow of PBS over the electrode. Record the current at the primary oxidation peak (~1.3 V) for each concentration to generate a calibration curve. - Selectivity Test: Repeat the injection procedure with potential interferents such as hydrogen peroxide (H₂O₂) and histamine to establish the unique voltammetric signature of adenosine at CNSMEs.
3. Data Analysis: - Calculate the sensitivity (nA/µM) from the slope of the adenosine calibration curve. Normalize this value by the electrode's capacitive background current to compare fairly with other electrode materials. CNSMEs typically show a 3- to 5-fold increase in normalized sensitivity for adenosine compared to bare CFMEs [17].
Protocol: Automated Adenosine Transient Detection using SSIM Image Analysis
This protocol describes the use of the Structural Similarity Index (SSIM) method for unbiased, accurate detection of adenosine release events in complex FSCV datasets [8].
1. Data Acquisition: - Collect FSCV data in vivo or in brain slices using standard CFMEs or CNSMEs and the FSCV waveform described in Protocol 4.1. Save data files in a compatible format (e.g., .mat or .txt for use in MATLAB).
2. Data Preprocessing: - High-Pass Filtering: Apply a 2nd-order Butterworth high-pass filter with a half-power frequency of 0.03 Hz to the raw FSCV data. This step removes low-frequency background drift and the large charging current, eliminating the need for traditional background subtraction [8]. - Data Formatting: Ensure the data is structured as a three-dimensional (3D) matrix: current (I) as a function of applied potential (E) and time (t), often visualized as a false-color plot.
3. SSIM Analysis Execution: - Software: Implement the SSIM algorithm in MATLAB 2019b or later. The code should compare sample data frames to a library of reference adenosine color plots. - Reference Library: Use a built-in standard library of 15 confirmed adenosine transient references, or manually identify 5-6 high-quality adenosine transients from the dataset to create an internal reference set. - Similarity Calculation: For each time point in the data stream, the algorithm calculates an SSIM index between the sample color plot and the reference plots. The SSIM index ranges from 0 (no similarity) to 1 (identical), accounting for luminance, contrast, and structure [8]. - Event Detection: Set an optimized SSIM index cutoff threshold (e.g., >0.7). Time points with SSIM indices exceeding this threshold are classified as adenosine release events.
4. Validation: - The performance of the SSIM method is quantified by its precision (fraction of detected events that are true positives, typically >99%) and recall (fraction of all true events detected, typically >95%), resulting in a high F1 score (>97%) [8].
Signaling Pathways and Experimental Workflows
Adenosine Detection Workflow
The diagram above illustrates the logical pathway from a physiological stimulus to the electrochemical detection of adenosine and its confirmed physiological role, providing an overview of the experimental process from cause to effect.
SSIM Analysis Process
The diagram above details the step-by-step data analysis pipeline using the Structural Similarity Index (SSIM), from raw data preprocessing to final event identification, which is critical for achieving high selectivity.
Advanced FSCV Methodologies for Sensitive and Selective Adenosine Sensing
Fast-scan cyclic voltammetry (FSCV) has emerged as a powerful technique for the real-time detection of neurochemical signaling, enabling researchers to monitor fluctuations in electroactive molecules with sub-second temporal resolution [10]. Within the broader thesis of advancing FSCV for adenosine detection research, this application note addresses a critical methodological focus: the optimization of key waveform parameters. Adenosine, a purine nucleoside with established roles as a neuromodulator and neuroprotector, regulates numerous physiological processes including sleep, blood flow, and neurotransmission [10] [11]. Unlike classical neurotransmitters, adenosine exhibits rapid, transient signaling events that last only seconds, necessitating detection methods capable of capturing these dynamics [10] [11] [20].
Traditional FSCV approaches, while successful for catecholamines, present specific challenges for adenosine detection due to its high oxidation potential and susceptibility to interference from compounds such as adenosine triphosphate (ATP) and hydrogen peroxide (H₂O₂) [10] [21]. The strategic selection of holding potentials, scan rates, and switching potentials is therefore paramount to enhancing sensitivity, selectivity, and temporal resolution for adenosine measurements. This protocol details optimized waveform parameters and methodologies that have been developed to address these challenges, providing researchers and drug development professionals with standardized approaches for characterizing adenosine signaling in both in vivo and in vitro preparations.
Waveform Parameters for Adenosine Detection
Fundamental Principles and Traditional Waveform
The detection of adenosine via FSCV relies on its electrochemical oxidation at carbon-fiber microelectrodes (CFMEs). Adenosine undergoes a series of irreversible oxidation steps, with the primary oxidation peak occurring at approximately 1.4 V and a secondary oxidation peak at 1.0 V versus a Ag/AgCl reference electrode [10]. The characteristic cyclic voltammogram (CV) featuring these two oxidation peaks serves as the electrochemical fingerprint for identifying adenosine [10] [20].
The traditional and most commonly applied waveform for adenosine detection is the triangular waveform. The specific parameters for this waveform, as established in the literature, are summarized in Table 1.
Table 1: Standard Triangular Waveform Parameters for Adenosine Detection
| Parameter | Value | Function/Rationale |
|---|---|---|
| Holding Potential | -0.4 V | Maintains a negative potential at the electrode between scans, which helps in attracting positively charged species and can enhance sensitivity for adenosine over ATP [10] [8]. |
| Switching Potential | 1.45 V to 1.50 V | Reaches the potential required to oxidize adenosine (primary peak at ~1.4 V). A higher potential ensures complete oxidation but may increase interference [10] [11] [20]. |
| Scan Rate | 400 V/s | Provides an optimal balance between current response (sensitivity) and the temporal resolution needed to capture rapid adenosine transients [10] [11]. |
| Repetition Rate | 10 Hz | Scans are repeated every 100 ms, enabling sub-second detection of adenosine dynamics [8] [11]. |
Advanced Waveform: The Sawhorse Optimization
To improve the selectivity of adenosine measurements against common interferents like ATP and H₂O₂, an advanced "sawhorse" waveform has been developed [21]. This modified waveform incorporates a brief holding period at the switching potential, which alters the resulting cyclic voltammograms of interferents, thereby facilitating better discrimination.
The key modification in the sawhorse waveform is a scan from -0.4 V to 1.35 V at 400 V/s, followed by a 1.0 ms hold at the switching potential, before ramping back down to -0.4 V [21]. The use of a slightly lower switching potential (1.35 V vs. 1.5 V) and the brief hold maximizes the time for adenosine oxidation while effectively altering the CV shapes of ATP and H₂O₂. Principal component analysis (PCA) has confirmed that the sawhorse waveform provides superior discrimination between adenosine, ATP, and H₂O₂ compared to the traditional triangle waveform [21].
Experimental Protocols
Electrode Preparation and Calibration
Carbon-Fiber Microelectrode (CFME) Fabrication:
- Materials: A single carbon fiber (e.g., T-650, 7 μm diameter) and a glass capillary for insulation [8] [11].
- Pulling: Aspirate the carbon fiber into the glass capillary and use a pipette puller to heat and pull the capillary, creating a sealed, tapered tip.
- Trimming: Carefully trim the exposed carbon fiber to a length of 65–100 μm beyond the glass seal [8] [11].
- Connection: Apply a conductive material (e.g., silver paint) to a lead wire and insert it into the glass capillary to establish an electrical connection with the carbon fiber [11].
Ag/AgCl Reference Electrode Preparation:
- Materials: A silver wire.
- Chloridization: Dip the tip of the silver wire in 1 M HCl and run a small electrical current through it for approximately 30 seconds to form a stable Ag/AgCl layer [11].
System Calibration:
- Flow Injection Analysis: Set up a flow cell system connected to a syringe pump and an injection valve.
- Standard Solutions: Prepare adenosine stock solutions (e.g., 10 mM in 0.1 M HClO₄) and dilute to working concentrations (e.g., 0.5 – 5 μM) in phosphate-buffered saline (PBS, pH 7.4) [11] [20].
- Data Collection: Flush the flow cell with buffer, inject standard adenosine solutions over the CFME, and record the FSCV data using the optimized waveform.
- Analysis: Plot the peak oxidation current against concentration to establish a calibration curve for converting nA signals to nM concentrations in vivo [11].
Data Acquisition and Analysis
In Vivo Measurement of Spontaneous Adenosine:
- Animal Preparation: Anesthetize the animal (e.g., rat) and secure it in a stereotaxic frame.
- Stereotaxic Implantation: Drill a burr hole and position the CFME in the brain region of interest (e.g., dorsal striatum or motor cortex). Implant the Ag/AgCl reference electrode in the contralateral hemisphere [11].
- Waveform Application: Apply the chosen waveform (triangular or sawhorse) continuously at 10 Hz. Allow the electrode to equilibrate for at least 30 minutes before beginning recordings to achieve a stable background current [11].
- Data Collection: Record electrochemical data continuously. Spontaneous, transient adenosine release events typically last for a few seconds and can be identified by their characteristic CV [11] [20].
Automated Data Analysis: The high data density of FSCV experiments (e.g., 144,000 CVs in a 4-hour recording) necessitates automated analysis [20].
- Background Subtraction: Subtract the stable background charging current to reveal Faradaic signals [10] [20].
- Algorithmic Detection: Use validated algorithms to identify adenosine transients. These algorithms typically exploit key features of adenosine:
- The presence of two oxidation peaks at the defined voltages.
- A time lag where the secondary peak follows the primary peak.
- A specific ratio of secondary-to-primary peak currents (typically between 0.49 and 0.89) [20].
- Advanced Image Analysis: For improved accuracy, employ image-based analysis such as the Structural Similarity Index (SSIM) method, which treats the entire 3D color plot as an image for pattern recognition. This method has demonstrated high precision (99.5%) and recall (95%) for detecting adenosine [8].
The Scientist's Toolkit
Table 2: Essential Research Reagents and Materials
| Item | Specification/Example | Primary Function |
|---|---|---|
| Carbon Fiber | T-650, 7 μm diameter (Cytec) [8] | The core sensing material of the microelectrode; provides a conductive, biocompatible surface for electron transfer during oxidation/reduction [10]. |
| Glass Capillaries | 1.2 mm x 0.68 mm (A-M Systems) [11] | Used to insulate and structurally support the carbon fiber, forming a sharp, sealed microelectrode tip. |
| Potentiostat | ChemClamp (Dagan) [8] | The core instrument that applies the precise voltage waveform to the working electrode and measures the resulting current. |
| Analysis Software | HDCV Analysis, Tar Heel CV, or custom MATLAB scripts [8] [11] [20] | Used for data acquisition, background subtraction, visualization via color plots, and quantitative analysis of neurotransmitter dynamics. |
| Adenosine Standard | >98% Purity (e.g., Acros Organics, Sigma-Aldrich) [11] [20] | For preparing calibration solutions to quantify electrode sensitivity and for use in in vitro pharmacological experiments. |
| Phosphate Buffered Saline (PBS) | pH 7.4, with added Ca²⁺/Mg²⁺ [8] | Provides a physiologically relevant ionic environment for in vitro calibrations and brain slice experiments. |
Visualization of Workflow and Waveforms
Experimental Workflow for Adenosine Detection
The following diagram outlines the comprehensive experimental pipeline for FSCV-based adenosine detection, from electrode preparation to data interpretation.
Figure 1: A flowchart detailing the key steps in an FSCV experiment for adenosine detection, highlighting critical phases of electrode preparation, waveform application, and data analysis.
Comparison of FSCV Waveforms for Adenosine
The core of this protocol lies in the selection and application of specific voltage waveforms. The diagram below illustrates the two primary waveforms discussed.
Figure 2: A comparison of the two primary FSCV waveforms used for adenosine detection, showing their parameters and specific use-case recommendations.
This application note has detailed the critical waveform parameters and experimental protocols for optimizing the detection of adenosine using FSCV. The precise configuration of holding potentials, scan rates, and switching potentials—whether using the standard triangular waveform or the advanced sawhorse variant—directly influences the sensitivity, selectivity, and overall success of adenosine measurements. By adhering to these standardized methodologies, researchers can reliably investigate the rapid dynamics of adenosine signaling, thereby advancing our understanding of its neuromodulatory and neuroprotective roles in health and disease. The integration of these electrochemical tools with other techniques, such as genetically encoded sensors, promises a future of rich, multi-modal analysis of neurochemical interactions [7].
Carbon microelectrodes (CMEs) are pivotal tools in neuroscience for the real-time monitoring of neurotransmitters, combining high temporal resolution with excellent biocompatibility. Their evolution from simple carbon fibers to sophisticated, nanomaterial-enhanced surfaces has significantly advanced our capacity to investigate complex neurochemical dynamics. This application note details the design principles, fabrication protocols, and performance characteristics of state-of-the-art CMEs. The content is framed within a research context focused on the detection of adenosine, a key neuromodulator, using fast-scan cyclic voltammetry (FSCV). Adenosine's role in processes like sleep-wake regulation and its neuroprotective effects, particularly its inhibition of dopamine and glutamate release via A1 receptors, makes it a critical target for electrochemical sensing [7] [22]. This guide is intended for researchers and scientists engaged in developing sensors for fundamental neuroscience and pharmaceutical applications.
Advanced Carbon Microelectrode Technologies
The pursuit of higher sensitivity, greater durability, and improved biocompatibility has driven the development of several advanced CME designs. The table below summarizes the key performance metrics of three prominent electrode types.
Table 1: Performance Comparison of Advanced Carbon Microelectrodes
| Electrode Type | Sensitivity (vs. 7µm CFME) | Key Advantage | Lifespan / Durability | In Vivo Dopamine Signal | Tissue Biocompatibility (Glial Activation) |
|---|---|---|---|---|---|
| 7 µm Bare CFME (Standard) | 1x (Baseline: 12.2 ± 4.9 pA/µm²) [9] | Minimal tissue damage [9] | Baseline | 24.6 ± 8.5 nA [9] | Baseline [9] |
| 30 µm Bare CFME | 2.7-fold higher in vitro [9] | Mechanical robustness & in vitro sensitivity [9] | - | 12.9 ± 8.1 nA (Reduced) [9] | Significantly higher [9] |
| 30 µm Cone-Shaped CFME | High (derived from improved signal) [9] | Optimized in vivo performance & biocompatibility [9] | 4.7-fold increase [9] | 47.5 ± 19.8 nA (3.7-fold improvement) [9] | Significantly lower (Iba1, GFAP) [9] |
| Carbon-Coated Microelectrode (CCM) | ~8-fold higher (125.5 nA/µM vs. 15.5 nA/µM for CFE) [23] | Scalability & integration with electrophysiology [23] | High electrochemical stability [23] | Validated in vivo (rodents) [23] | - |
Cone-Shaped Carbon Fiber Microelectrodes (CFMEs)
Increasing the diameter of carbon fibers from the standard 7 µm to 30 µm enhances mechanical strength and in vitro sensitivity. However, this comes at the cost of increased tissue damage and compromised in vivo signal quality upon implantation. To mitigate this, a cone-shaped geometry, achieved via electrochemical etching, has been developed. This design ensures a sharper tip, which facilitates smoother tissue penetration, minimizes insertion-induced damage, and results in significantly lower glial activation (as measured by Iba1 and GFAP markers) alongside a 3.7-fold improvement in in vivo dopamine signals and a 4.7-fold increase in operational lifespan compared to standard 7 µm CFMEs [9].
Carbon-Coated Microelectrodes (CCMs)
A transformative approach involves coating conventional gold microelectrodes with a graphene-based carbon layer. This process involves electroplating via potentiostatic deposition of graphene oxide (GO) followed by mild annealing at 250 °C in an N₂ environment. This annealing step is crucial, as it drastically improves electrochemical stability by reducing the interlayer spacing from 4.0 Å to 3.7 Å and lowering the oxygen content from 15.9% to 8.7%, thereby resisting water/ion infiltration [23]. The CCMs exhibit exceptional performance, with a dopamine sensitivity of 125.5 nA/µM and a low limit of detection of 5 nM. A key advantage of this technology is its scalability and compatibility with standard microfabrication processes, enabling the creation of high-density arrays (e.g., 100 channels) and seamless integration with electrophysiological recording sites for dual-mode sensing [23].
Experimental Protocols
Protocol: Fabrication of 30 µm Cone-Shaped CFMEs
This protocol details the creation of cone-shaped CFMEs for improved chronic implantation [9].
Key Reagents & Equipment:
- 30 µm diameter carbon fiber (World Precision Instruments).
- Tris buffer (15 mM Trizma phosphate, 3.25 mM KCl, 140 mM NaCl, 1.2 mM CaCl₂, 1.25 mM NaH₂PO₄, 1.2 mM MgCl₂, and 2.0 mM Na₂SO₄, pH 7.4).
- Homemade electrochemical etching system with a linear actuator.
- Capillary puller and scalpel.
Procedure:
- Electrode Fabrication: Aspirate a single 30 µm carbon fiber into a glass capillary and pull it to a microscopic tip using a pipette puller. Trim the exposed fiber to approximately 100 µm in length using a scalpel [9].
- Electrochemical Etching:
- Submerge a 1 mm segment of the carbon fiber in Tris buffer.
- Apply a direct current voltage of 10 V for 20 seconds to initiate electrolysis and partial erosion of the fiber.
- Simultaneously, actuate a linear actuator to move the electrode upward at a constant speed. This controlled withdrawal from the solution is what forms the cone shape.
- Adjust the actuator speed to control the final cone height, typically between 100 and 120 µm [9].
- Pre-conditioning: Before detection, precondition the CFME using a 1.5 V FSCV sweep (−0.4 to 1.5 V at 400 V/s, 30 Hz), followed by application of the standard FSCV waveform (−0.4 to 1.3 V sweep at 10 Hz) [9].
Protocol: FSCV for Adenosine and Dopamine Comonitoring
This protocol describes the use of FSCV for the simultaneous, real-time detection of adenosine and dopamine, relevant for studying neuromodulation in systems like deep brain stimulation [22].
Key Reagents & Equipment:
- Polyacrylonitrile-based carbon fiber (T-650, 5-µm diameter).
- WINCS (Wireless Instantaneous Neurotransmitter Concentration System) or equivalent FSCV potentiostat.
- Ag/AgCl reference electrode.
- Tris-buffered saline (150 mM NaCl, 12 mM Tris-base, pH 7.4).
Procedure:
- FSCV Parameters:
- Use a triangular waveform, scanning from −0.4 V to +1.5 V and back.
- Apply a scan rate of 400 V/s at a frequency of 10 Hz (every 0.1 seconds).
- Hold the working electrode at a bias potential of −0.4 V between scans [22].
- Flow Injection Analysis (for in vitro calibration):
- Place the CFME in a stream of Tris buffer flowing at 2 mL/min.
- Inject a 1 mL bolus of analyte (adenosine, dopamine, or a combination) into the stream for 5-10 seconds.
- Identify analytes based on their characteristic peak potentials: adenosine oxidizes at +1.5 V and +1.0 V, while dopamine oxidizes at +0.6 V [22].
- In Vivo Measurement:
- Implant the CFME in the target brain region (e.g., caudate putamen).
- Place an Ag/AgCl reference electrode in superficial cortical tissue.
- Use electrical stimulation (e.g., of the ventral tegmental area/substantia nigra) to evoke neurotransmitter release.
- The WINCS system wirelessly transmits the FSCV data to a base station for analysis with software such as MATLAB or LabVIEW [22].
- FSCV Parameters:
Protocol: Multiplexing FSCV with Fluorescence Sensors
This advanced protocol allows for the simultaneous monitoring of electroactive (e.g., dopamine, adenosine) and non-electroactive (e.g., glutamate) neurotransmitters [7].
Key Reagents & Equipment:
- Sindbis viral vector for expression of iGluSnFR3.v857 (genetically encoded glutamate sensor).
- Standard FSCV setup with CFME.
- Fluorescence microscopy setup.
Procedure:
- Sensor Expression: Express the iGluSnFR3.v857 glutamate sensor in the target brain region (e.g., caudate-putamen) of a brain slice using a Sindbis viral vector. Expression typically requires 18-24 hours [7].
- Simultaneous Recording:
- Implant a CFME into the brain slice near cells expressing the glutamate sensor.
- Use the CFME with FSCV to monitor electrically stimulated dopamine (and/or adenosine) release.
- Simultaneously, use fluorescence microscopy to record changes in glutamate signaling via the iGluSnFR3 sensor.
- Pharmacological Manipulation: To investigate receptor-specific mechanisms (e.g., adenosine's action via A1 receptors), locally apply agents like an A1 receptor antagonist (e.g., DPCPX) and observe the effects on the recorded signals [7].
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents and Materials for Carbon Microelectrode Research
| Item | Function / Application | Example / Specification |
|---|---|---|
| Carbon Fibers | The core sensing material for CFMEs. Choice affects electron transfer kinetics and background current. | Polyacrylonitrile (PAN)-based (e.g., T-650 for fast kinetics) or Pitch-based (e.g., P-55 for high current) [18]. |
| Graphene Oxide Dispersion | Precursor for electroplating carbon coatings on gold microelectrodes to create CCMs. | Aqueous dispersion for potentiostatic deposition [23]. |
| Tris Buffer | Electrochemically stable buffer for in vitro calibration and electrochemical etching. | 15 mM Trizma phosphate, 140 mM NaCl, pH 7.4 [9]. |
| Adenosine & Dopamine Stock Solutions | Primary analytes for calibration and in vivo detection. | Dissolve adenosine in pure water; dissolve dopamine in 1 mM perchloric acid to prevent oxidation [22]. |
| iGluSnFR3.v857 | Genetically encoded fluorescence sensor for multiplexed detection of non-electroactive glutamate. | Expressed via Sindbis viral vector in target brain tissue [7]. |
| DPCPX (8-Cyclopentyl-1,3-dipropylxanthine) | Selective A1 receptor antagonist for probing adenosine's mechanistic role in neuromodulation [7]. | - |
| Ag/AgCl Reference Electrode | Essential stable reference for all electrochemical measurements. | Chloridized silver wire [22]. |
Signaling Pathway and Experimental Logic
Understanding the neurochemical interplay is crucial for designing relevant experiments. Adenosine, released during neural activity or pathological events, exerts its effects primarily through inhibitory A1 receptors.
This diagram illustrates the central role of A1 receptors in mediating adenosine's inhibitory effects on other neurotransmitters, a key pathway for investigation using the protocols described.
The detection of rapid neurochemical signaling, particularly of neuromodulators like adenosine, is crucial for understanding brain function and developing new therapeutic strategies. Fast-scan cyclic voltammetry (FSCV) has emerged as a primary technique for monitoring these dynamics with sub-second temporal resolution [4] [11]. However, a significant challenge has been the accurate and automated identification of analytes, especially in complex mixtures or when signals exhibit evolving electrochemical signatures [8]. Traditional analysis methods, which often focus on single cyclic voltammograms or specific current-time traces, fail to fully leverage the rich, three-dimensional data structure of FSCV experiments (current vs. potential vs. time) [8] [24].
The integration of image analysis techniques, specifically the Structural Similarity Index (SSIM), represents a transformative approach for automated detection in electrochemical data. SSIM is a perceptual metric that quantifies the similarity between two images based on comparisons of luminance, contrast, and structure [25] [26]. Originally developed for assessing digital image and video quality, its application has expanded to fields like mass spectrometry and, most recently, electrochemistry [8] [27]. By treating FSCV color plots as images, the SSIM algorithm can perform a sophisticated, holistic comparison of sample data against reference signals, enabling high-fidelity detection of neurotransmitters and neuromodulators in an automated, efficient manner [8]. This protocol details the application of SSIM-based analysis for the detection of adenosine using FSCV.
The SSIM Algorithm: Principles and Application to FSCV
Core Mathematical Principles
The Structural Similarity Index (SSIM) is a full-reference metric, meaning it assesses the quality or similarity of a test image against a reference image. The algorithm is based on the human visual system's sensitivity to structural information and is calculated using local patterns of pixel intensities normalized for luminance and contrast [25] [28].
The general formula for SSIM is a combination of three comparative components: luminance ((l)), contrast ((c)), and structure ((s)):
[ SSIM(x, y) = [l(x, y)]^{\alpha} \cdot [c(x, y)]^{\beta} \cdot [s(x, y)]^{\gamma} ]
Where:
- (x) and (y) represent the two image windows being compared.
- (\alpha), (\beta), and (\gamma) are parameters to adjust the relative importance of the three components [25].
In its most common and simplified form, with (\alpha = \beta = \gamma = 1) and (c3 = c2/2), the formula reduces to:
[ SSIM(x, y) = \frac{(2\mux\muy + c1)(2\sigma{xy} + c2)}{(\mux^2 + \muy^2 + c1)(\sigmax^2 + \sigmay^2 + c_2)} ]
With:
- (\mux) and (\muy) representing the local pixel sample means (luminance).
- (\sigmax) and (\sigmay) representing the standard deviations (contrast).
- (\sigma_{xy}) representing the cross-covariance between (x) and (y) (structure).
- (c1) and (c2) being stabilizing constants (c1 = (k1L)^2), (c2 = (k2L)^2), where (L) is the dynamic range of pixel values and (k1 \ll 1), (k2 \ll 1) are small constants (typically (k1=0.01), (k2=0.03)) [25].
The SSIM index yields a value between -1 and 1, where 1 indicates perfect similarity, 0 indicates no similarity, and -1 indicates perfect anti-correlation [25] [26].
SSIM as an Image Recognition Tool for FSCV
In FSCV, data is visually represented as a color plot, a three-dimensional image where the x-axis represents time, the y-axis represents the applied potential, and the color represents the measured current. Each electroactive species generates a unique "fingerprint" in this color plot based on its distinct redox properties [8].
Applying SSIM to FSCV involves treating transient neurochemical events in the color plot as images to be recognized. The process involves:
- Creating a Reference Library: Collecting canonical FSCV color plot "images" for the target analyte (e.g., adenosine) from validated in vitro or in vivo recordings.
- Sliding Window Analysis: A sample window (e.g., a few seconds of FSCV data) is slid across the entire dataset.
- Similarity Calculation: For each position of the sample window, the SSIM index is calculated against the pre-defined reference images.
- Event Detection: A high SSIM score (close to 1) indicates a strong structural similarity between the sample data and the reference, signaling a detection event [8].
This method outperforms traditional peak-based analyses because it considers the entire spatial structure of the voltammetric data simultaneously, making it more robust to noise and capable of identifying analytes with complex, time-varying cyclic voltammograms, such as adenosine [8].
Application Notes: SSIM for Adenosine Detection
Experimental Setup and Reagent Solutions
The following table details the essential materials and reagents required for conducting FSCV experiments for adenosine detection and subsequent SSIM analysis.
Table 1: Key Research Reagents and Materials for FSCV Adenosine Detection
| Item | Function / Description | Example Details / Source |
|---|---|---|
| Carbon-Fiber Microelectrode (CFME) | Working electrode for FSCV measurements. T-650 carbon fibers (7-μm diameter) are sealed in a glass capillary with ~100 μm exposed [8] [11]. | Cytec Engineering Materials [8]. |
| Ag/AgCl Reference Electrode | Provides a stable reference potential for the electrochemical cell [8]. | |
| Adenosine | Primary analyte of interest. Prepared as a stock solution in 0.1 M HClO4 and diluted in PBS for working standards [8]. | Acros Organics [8]. |
| Phosphate Buffered Saline (PBS) | Physiological buffer for in vitro calibrations and experiments. Mimics the ionic composition of the extracellular brain environment [8]. | 131.25 mM NaCl, 3.00 mM KCl, 10.0 mM NaH2PO4, 1.2 mM MgCl2, 2.0 mM Na2SO4, 1.2 mM CaCl2, pH 7.4 [8]. |
| FSCV Potentiostat | Instrumentation to apply the waveform and measure the resulting current. | e.g., ChemClamp potentiostat (Dagan) [8]. |
| Data Acquisition Software | Software to control the potentiostat and collect the raw current-time data. | e.g., HDCV Analysis (University of North Carolina) [8]. |
| MATLAB with Custom Scripts | Software environment for implementing digital filtering, SSIM calculation, and automated detection algorithms [8]. | MathWorks [8]. |
Critical Experimental Parameters for Adenosine
Adenosine presents a unique detection challenge because it undergoes multiple oxidation steps, resulting in a cyclic voltammogram with a primary oxidation peak at approximately 1.3 V and a secondary oxidation peak that grows in over time [8] [24]. This temporal evolution of the CV shape makes it difficult for traditional analysis techniques but is well-suited for SSIM, which can account for these structural changes across the entire color plot.
The typical FSCV waveform parameters optimized for adenosine detection are:
- Holding Potential: -0.4 V (vs. Ag/AgCl)
- Switching Potential: +1.45 V
- Scan Rate: 400 V/s
- Repetition Rate: 10 Hz (every 100 ms) [8] [11]
SSIM Performance in Adenosine Detection
The implementation of SSIM analysis for spontaneous adenosine release has demonstrated exceptional performance, outperforming previous algorithms that relied on current-time traces.
Table 2: Quantitative Performance Metrics of SSIM for Adenosine Detection
| Performance Metric | SSIM Method Result (Mean ± SD) | Description and Implication |
|---|---|---|
| Precision | 99.5 ± 0.6% | The proportion of detected events that are true positives. Very high precision indicates minimal false positives. |
| Recall (Sensitivity) | 95 ± 3% | The proportion of true events that are successfully detected. High recall indicates the algorithm misses very few real events. |
| F1 Score | 97 ± 2% | The harmonic mean of precision and recall. A single metric summarizing overall detection accuracy (closer to 100% is better). |
| Selectivity | Successfully rejected interferents | The method effectively distinguished adenosine from common interferents like pH changes, histamine, and H2O2 [8]. |
This data is based on 15 experiments from three different researchers, confirming the robustness and reliability of the method [8].
Detailed Protocols
Protocol 1: SSIM-Based Automated Detection of Adenosine Transients
This protocol describes the step-by-step procedure for analyzing FSCV data to detect rapid adenosine release events using the Structural Similarity Index.
I. Materials and Software
- Raw FSCV data files (e.g., in a compatible format for MATLAB).
- MATLAB 2019b or later with Image Processing Toolbox.
- Custom SSIM analysis software (e.g., as described in [8]).
- Pre-established reference color plots for adenosine.
II. Procedure
- Data Preprocessing:
- Apply a high-pass, second-order Butterworth filter to the raw FSCV current data. Use a half-power frequency of 0.03 Hz to remove background drift and low-frequency noise [8].
- (Optional) For additional noise reduction, apply a Savitzky-Golay filter (e.g., window length of 15) to smooth the data without significantly distorting the signal [8].
Reference Selection ("Internal Reference" Method):
- From the dataset to be analyzed, manually identify the start time of six clear, representative transient adenosine events.
- Extract these events to serve as the internal reference library for that specific dataset [8].
- Alternative: Use a "Standard Library" of 15 adenosine references built into the software, which may be more generalizable.
SSIM Calculation and Event Detection:
- Normalize both the sample data windows and the reference color plots to their respective maximum currents.
- For each time point in the sample data, extract a window of the color plot centered on that time.
- Calculate the SSIM index between the sample window and each reference in the library using the formula in Section 2.1.
- The final SSIM score for the sample window is the highest score obtained from all comparisons.
- Compare the final SSIM score against a pre-optimized cutoff threshold. Scores above the threshold are classified as adenosine events.
Validation and Output:
- The algorithm outputs a list of detected event times.
- Compare the results against a benchmark method (e.g., the "Borman Method" [8]) or manual scoring by an expert to calculate precision, recall, and F1 score (as in Table 2).
Protocol 2: In Vitro Calibration and Selectivity Testing
This protocol ensures the carbon-fiber microelectrode is sensitive and selective to adenosine before in vivo application.
I. Materials
- Calibration setup with flow cell and syringe pump.
- Stock solutions of adenosine, dopamine, histamine, and H2O2.
- PBS buffer, pH 7.4.
II. Procedure
- Electrode Calibration:
- Place the CFME in the flow cell with a constant flow of PBS.
- Using a loop injector, inject known concentrations of adenosine (e.g., 0.5, 1.0, 5.0 μM) into the flow stream.
- Record the FSCV response at each concentration.
- Plot the peak oxidation current versus concentration to generate a calibration curve and determine the electrode's sensitivity (nA/μM) and limit of detection (LOD).
- Selectivity Testing:
- In the flow cell, sequentially inject solutions of potential interferents (e.g., pH changes from 7.3 to 7.5, 1-10 μM histamine, 1-10 μM H2O2) and record the FSCV responses.
- Collect the resulting color plots for each interferent to be used as negative references.
- Process the data using the SSIM algorithm optimized for adenosine. Confirm that the SSIM scores for interferent signals fall below the detection threshold, demonstrating selectivity [8].
Advanced Applications and Broader Context
The SSIM method is a generalizable strategy that extends beyond adenosine. It has been successfully optimized for the detection of dopamine and, importantly, can resolve simultaneous adenosine and dopamine release events, which is a significant challenge for traditional analyses [8]. This highlights the power of image-based analysis to deconvolve complex neurochemical mixtures.
The integration of automated detection algorithms like SSIM is part of a broader trend in biosensing and medical diagnostics towards leveraging artificial intelligence (AI) and machine learning (ML). These tools are critical for processing the high-dimensional data generated by modern sensors, converting complex signals into actionable, clinically relevant information [29] [30]. The SSIM-based approach for FSCV is a prime example of how modern data analysis techniques can overcome the limitations of classical methods, paving the way for more sophisticated, efficient, and automated analysis in neuroscience research and drug development.
Multiplexing FSCV with Fluorescent Sensors for Simultaneous Neurotransmitter Measurement
Understanding the complex interactions between different neurotransmitters is fundamental to deciphering brain function and developing treatments for neurological disorders. A significant challenge in neuroscience has been the simultaneous measurement of multiple neurotransmitters, which often have intertwined and dynamic relationships. Fast-scan cyclic voltammetry (FSCV) is a powerful electrochemical technique known for its high temporal resolution (sub-second) and excellent sensitivity for detecting electroactive analytes like dopamine and adenosine [7] [4]. However, its utility is limited for nonelectroactive neurotransmitters, such as glutamate. Conversely, genetically encoded fluorescent sensors offer high spatial resolution and specificity for a wide range of neurochemicals, including nonelectroactive ones, but can be limited by photobleaching and do not provide direct concentration information [7].
This application note details a novel methodology that multiplexes FSCV with fluorescent sensors to overcome the limitations of either technique used in isolation. We focus on the simultaneous detection of adenosine, dopamine, and glutamate to investigate the rapid neuromodulatory role of adenosine in the brain, a key theme in advanced FSCV research [7] [4]. This protocol provides a framework for researchers to capture complementary chemical information in real-time, enabling a more holistic view of neurochemical signaling.
Experimental Principles and Workflow
The core principle of this multiplexed approach is the synergistic combination of FSCV's high temporal resolution for electroactive neurotransmitters with the high spatial and molecular specificity of genetically encoded fluorescent sensors for nonelectroactive neurotransmitters [7]. The experimental workflow can be visualized as follows:
This integrated approach reveals interactions that would be missed by single-technique measurements. For instance, it has been used to demonstrate that adenosine exerts a transient inhibitory effect on both dopamine and glutamate release via A1 receptors, with effects spatially constrained within a 250 μm radius [7].
Key Research Reagent Solutions
The successful implementation of this multiplexed technique relies on a set of specific reagents and tools. The table below catalogues the essential components.
Table 1: Essential Research Reagents and Materials
| Item | Function/Description | Key Details |
|---|---|---|
| iGluSnFR3.v857 | Genetically encoded glutamate sensor | High sensitivity, fast binding kinetics; expressed via Sindbis viral vector [7]. |
| Carbon Fiber Microelectrode (CFME) | Working electrode for FSCV detection | Detects electroactive analytes (DA, AD); 7-30 μm diameter [7] [9]. |
| A1 Receptor Antagonist (DPCPX) | Pharmacological tool for receptor mechanism validation | Blocks A1 receptors to confirm adenosine's action pathway [7]. |
| HDCV Software | Data acquisition and analysis for FSCV | Enables simultaneous data collection from multiple electrodes and integration with behavioral or digital inputs [31]. |
| Sindbis Viral Vector | Vehicle for rapid sensor expression | Enables high-level sensor expression in mammalian cells within 18-24 hours [7]. |
Detailed Experimental Protocols
Sensor Expression and Brain Slice Preparation
This initial phase prepares the biological substrate for the multiplexed measurements.
- Viral Injection: Sterotaxically inject a Sindbis viral vector carrying the iGluSnFR3.v857 gene into the target brain region (e.g., caudate-putamen) of anesthetized mice [7].
- Incubation: Allow 18-24 hours for robust sensor expression in the target region [7].
- Slice Preparation: Euthanize the animal and prepare acute coronal brain slices (300-400 μm thick) containing the transfected region in ice-cold, oxygenated (95% O₂/5% CO₂) artificial cerebrospinal fluid (aCSF).
- Recovery & Verification: Incubate slices for at least 1 hour in aCSF at room temperature for recovery. Verify sensor expression and localization using fluorescence microscopy.
Multiplexed FSCV and Fluorescence Recording
This core protocol describes the setup and execution of the simultaneous measurement.
Electrochemical Setup:
- Fabricate a CFME and precondition it using a standard FSCV waveform (e.g., -0.4 V to +1.3 V and back, 400 V/s, 10 Hz) until a stable background current is achieved [9].
- Position the CFME in the brain slice, within 250 μm of cells expressing iGluSnFR3.v857 [7].
- Use the HDCV data acquisition system or similar to apply the FSCV waveform and record the faradaic current [31].
Optical Setup:
- Position the brain slice under an epifluorescence or conforescence microscope.
- Set the appropriate excitation wavelength for iGluSnFR3.v857 and collect emitted light through a bandpass filter.
- Use a high-sensitivity camera (e.g., sCMOS) to record fluorescence changes at a high frame rate (≥10 Hz).
Stimulation and Pharmacological Application:
- Electrical Stimulation: Place a bipolar stimulating electrode upstream of the recording site. Apply a monophasic, constant-current pulse train to evoke endogenous neurotransmitter release.
- Adenosine Application: Apply exogenous adenosine locally to the slice via a micropipette for a short duration (e.g., 30 seconds) [7].
- Receptor Blockade: To confirm the role of A1 receptors, pre-apply and continuously perfuse the selective antagonist DPCPX (e.g., 100 nM) into the aCSF [7].
Data Integration and Analysis
The final phase involves processing and correlating the data streams.
FSCV Data:
- Process the recorded currents using HDCV Analysis or similar software. This includes background subtraction, digital filtering (e.g., 2D FFT), and conversion to chemical information [31].
- Identify analytes (dopamine, adenosine) by their characteristic cyclic voltammogram peaks.
- Use principal component regression (PCR) to extract concentration-time profiles for multiple overlapping analytes [31].
Fluorescence Data:
- Process the video data to extract fluorescence intensity (ΔF/F) over time from regions of interest (ROIs) corresponding to the sensor expression sites.
- The fluorescence signal is a proxy for glutamate concentration but requires calibration for quantitative analysis.
Data Correlation:
- Temporally align the processed FSCV concentration traces and fluorescence traces using a common timing signal recorded by the acquisition software [31].
- Analyze the relationships between the different neurotransmitters, such as the correlation between glutamate and dopamine release and their coordinated response to adenosine application.
Quantitative Data and Key Findings
The application of this multiplexed approach yields robust quantitative data on neurotransmitter dynamics and interactions.
Table 2: Quantitative Summary of Adenosine's Inhibitory Effects
| Parameter | Dopamine Release | Glutamate Release | Measurement Technique |
|---|---|---|---|
| Inhibition by Adenosine | Transient Inhibition | Transient Inhibition | FSCV & Fluorescence [7] |
| Spatial Range of Effect | Within 250 μm | Within 250 μm | Spatial profiling [7] |
| Recovery Time | ~10 minutes | ~10 minutes | Post-application recording [7] |
| Receptor Mechanism | A1 Receptor | A1 Receptor | Blocked by DPCPX [7] |
| Spatial Correlation | Inverse correlation with glutamate release | Inverse correlation with dopamine release | Simultaneous measurement [7] |
The signaling pathway elucidated by these experiments, which can be inhibited by DPCPX, is summarized below.
The multiplexing of FSCV with genetically encoded fluorescent sensors represents a significant technological advancement for the real-time monitoring of complex neurochemical signaling. This protocol provides a detailed guide for simultaneously tracking adenosine, dopamine, and glutamate, revealing rapid, transient, and spatially confined neuromodulatory interactions that are central to a comprehensive thesis on adenosine research. By integrating the high temporal resolution of electrochemistry with the molecular and spatial specificity of optical sensors, this approach opens new avenues for investigating the intricate chemical conversations that underlie brain function and dysfunction.
Overcoming Technical Hurdles: Enhancing FSCV Performance and Durability
Strategies for Mitigating Electrode Biofouling and Surface Passivation
Electrode biofouling and surface passivation represent significant challenges in fast-scan cyclic voltammetry (FSCV) for adenosine detection, particularly during prolonged in vivo measurements. Biofouling refers to the accumulation of biomolecules (e.g., proteins, lipids) on electrode surfaces, while surface passivation encompasses the chemical degradation of electrode materials through processes such as the deposition of oxidative by-products or exposure to interfering ions [32]. In FSCV, these phenomena manifest as a shift in the peak oxidative potential of the background signal (Eₚ,ᵦₖ𝓰) and diminished sensitivity to target analytes like adenosine [33]. The immune response to chronically implanted electrodes triggers a cascade of events that collectively degrade voltammetric performance, ultimately compromising the accuracy and reliability of adenosine measurements in long-term studies [33] [32]. This application note details targeted strategies to mitigate these effects, ensuring data integrity throughout experimental timelines.
Underlying Mechanisms of Electrode Fouling
Primary Fouling Pathways
Understanding the distinct mechanisms of electrode fouling is essential for developing effective mitigation strategies. Biofouling and chemical fouling affect electrode systems through different pathways, each requiring specific countermeasures.
Table 1: Characteristics of Electrode Fouling Mechanisms
| Fouling Mechanism | Primary Causes | Effects on Working Electrode (CFME) | Effects on Reference Electrode (Ag/AgCl) |
|---|---|---|---|
| Biofouling | Accumulation of proteins (e.g., BSA), lipids, and other biomolecules [32] | Decreased sensitivity, peak voltage shifts, reduced electron transfer kinetics [32] | Minimal direct impact on FSCV signals [32] |
| Chemical Fouling | Adsorption of oxidative by-products from neurotransmitters (e.g., serotonin, dopamine) [32] | Significant sensitivity loss, altered background current, fouling-induced peak shifts [32] | Not directly applicable |
| Reference Electrode Fouling | Exposure to sulfide (S²⁻) ions in biological environments [32] | Not applicable | Cathodic polarization, decreased open circuit potential (OCP), peak potential shifts in FSCV [33] [32] |
The experimental workflow for identifying and addressing these fouling mechanisms can be visualized as follows:
Impact on Adenosine Detection
For adenosine detection specifically, fouling mechanisms present unique challenges. Adenosine detection using FSCV relies on maintaining stable electron transfer kinetics and surface adsorption properties, both of which are compromised by fouling. The similarity in oxidation potentials between adenosine and interferents like hydrogen peroxide makes fouling-induced peak shifts particularly problematic for accurate identification and quantification [34]. Furthermore, the low concentrations of extracellular adenosine in physiological systems necessitate high sensitivity, which is dramatically reduced by both biofouling and chemical fouling mechanisms [35] [34].
Mitigation Strategies and Experimental Protocols
Electrode Configuration and Design
Three-Electrode Configuration for Long-Term Implantation The conventional two-electrode configuration for FSCV becomes unsuitable for long-term studies due to increased electrochemical impedance from electrode encapsulation. Implementing a three-electrode system with a dedicated counter electrode compensates for this increased impedance and reduces the Eₚ,ᵦₖ𝓰 shift in vivo [33].
Protocol 3.1.1: Implementing Three-Electrode Configuration
- Electrode Preparation: Fabricate carbon-fiber working electrodes as described in Protocol 3.2.1. Prepare Ag/AgCl reference electrodes by chloridizing silver wire in sodium hypochlorite for 24 hours. Use platinum wire (0.25 mm diameter) as the counter electrode [33].
- Surgical Implantation: Lower the carbon-fiber electrode into the target brain region (e.g., nucleus accumbens). Implant the Ag/AgCl reference electrode and Pt-wire counter electrode in the contralateral hemisphere, approximately 4 mm into brain tissue [33].
- Headstage Connection: Utilize a headstage designed specifically for three-electrode configuration to properly compensate for impedance components of biofouling [33].
- Validation: Confirm system performance by measuring dopamine sensitivity at artificially increased impedance levels in vitro before proceeding with in vivo adenosine measurements [33].
Electrode Material Selection and Fabrication
Carbon-Fiber Microelectrode Fabrication with Enhanced Materials The choice of electrode materials significantly impacts fouling resistance. Boron-doped diamond microelectrodes (BDDMEs) demonstrate superior resistance to biofouling compared to traditional carbon-fiber microelectrodes (CFMEs), particularly for challenging analytes like serotonin [36].
Protocol 3.2.1: CFME Fabrication for Adenosine Detection
- Fiber Preparation: Insert a single 7.1 μm-diameter carbon fiber into a 15 mm-long fused silica capillary (20-μm ID, 90-μm OD) in isopropanol [33].
- Sealing: Seal one end of the capillary with Devcon two-ton epoxy, leaving a small length of exposed carbon fiber [33].
- Electrical Connection: On the opposite end, connect the carbon fiber to a gold-plated beryllium-copper-nickel pin with alcohol-based graphite conductive adhesive, followed by epoxy application [33].
- Trimming: Trim the exposed carbon fiber to a length of 100 μm with a scalpel under a brightfield microscope [33].
- Optional Surface Modification: For enhanced adenosine selectivity, modify carbon paste electrodes with porous materials such as natural phosphate, which strongly adsorbs adenosine [35].
Boron-Doped Diamond Microelectrodes for Fouling Resistance BDDMEs exhibit inherent resistance to fouling due to their sp³ carbon structure, extended π-electron system, and fewer carbon-oxygen surface groups, which reduce molecular adsorption [36].
Protocol 3.2.2: BDDME Implementation for Fouling-Prone Environments
- Electrode Selection: Utilize wafer-fabricated, freestanding all-diamond BDDMEs with polycrystalline diamond insulation [36].
- Waveform Optimization: Employ the "Jackson" waveform (0.2 V to 1.0 V to -0.1 V to 0.2 V at 1000 V s⁻¹) for optimal biofouling performance with BDDMEs [36].
- Performance Validation: Compare 5-HT responses at BDDMEs and CFMEs under biofouling conditions (e.g., in protein solutions) to confirm superior fouling resistance [36].
Surface Modifications and Coatings
Antifouling Coatings for Working Electrodes Surface coatings create a physical barrier that prevents fouling agents from reaching the electrode surface while maintaining analyte permeability.
Table 2: Antifouling Coating Strategies for Electrodes
| Coating Type | Composition | Application Method | Fouling Reduction Efficacy | Compatibility with Adenosine Detection |
|---|---|---|---|---|
| Nafion Coating | Cation-exchange polymer | Dip-coating or electrochemical deposition | Moderate biofouling reduction [33] | Compatible; may affect adenosine adsorption |
| PEDOT:Nafion | Conductive polymer composite | Electrochemical polymerization | Dramatic reduction in acute in vivo biofouling [32] | Requires validation for adenosine |
| PEDOT-PC | Phosphorylcholine functionalized ethylene-dioxythiophene | Electrochemical polymerization | Significant reduction in biomacromolecule accumulation [32] | Promising for in vivo adenosine detection |
| NanoMIPs | Molecularly imprinted polymers with modified thymidine monomers | Solid-phase synthesis with glutaraldehyde linker | High selectivity for adenosine (K_D = 2.11 nM) [37] | Excellent for specific adenosine detection |
Protocol 3.3.1: Application of PEDOT:PSS Antifouling Coating
- Solution Preparation: Prepare a solution of 0.1 M PEDOT:PSS in phosphate buffer (pH 7.4) [32].
- Electrode Pretreatment: Clean CFMEs by applying extended triangular waveform (-0.4 V to +1.4 V at 400 V s⁻¹) in aCSF for 30 minutes [34].
- Electrodeposition: Immerse the CFME in PEDOT:PSS solution and apply constant potential of +1.0 V vs. Ag/AgCl for 30 seconds [32].
- Curing: Rinse gently with DI water and air-dry for 1 hour before use [32].
- Validation: Test coated electrodes in BSA solution (40 g L⁻¹) for 2 hours while applying FSCV waveform to confirm fouling resistance [32].
Reference Electrode Protection Reference electrode fouling primarily occurs through sulfide ion exposure, leading to cathodic polarization and OCP decrease [32].
Protocol 3.3.2: Nafion-Coating Ag/AgCl Reference Electrodes
- Chloridization: Soak silver wire (0.25 mm diameter) in 8.25% sodium hypochlorite for 24 hours to form Ag/AgCl layer [33].
- Nafion Preparation: Prepare 5% Nafion solution in appropriate solvent [33].
- Coating Application: Dip the chloridized Ag/AgCl wire in Nafion solution and withdraw at controlled rate (1 mm/s) [33].
- Drying: Cure at 70°C for 10 minutes to form protective membrane [33].
- Performance Testing: Measure OCP in buffer with and without sulfide ions (1 mM) to confirm delayed polarization onset [33] [32].
Waveform Optimization Strategies
Extended Waveform Parameters for Fouling Mitigation Strategic adjustment of FSCV waveform parameters can reduce fouling and maintain electrode sensitivity.
Protocol 3.4.1: Waveform Optimization for Fouling Resistance
- Switching Potential Extension: Extend switching potential from +1.0 V to +1.4 V to enhance sensitivity and promote surface cleaning through breakage of carbon-carbon bonds and addition of edge plane sites [34] [36].
- Holding Potential Adjustment: Utilize more negative holding potentials (-0.6 V instead of -0.4 V) to enhance adsorption of cationic molecules, improving sensitivity for adenosine detection [34].
- Waveform Selection: For serotonin-rich environments where chemical fouling is prevalent, implement the "Jackson" waveform (0.2 V to 1.0 V to -0.1 V at 1000 V s⁻¹) to minimize fouling while maintaining detection capability [36].
- Frequency Optimization: Adjust repetition frequency based on temporal resolution requirements, noting that BDDMEs show more sustained responses to frequency changes than CFMEs [36].
The Scientist's Toolkit: Essential Research Reagents and Materials
Table 3: Key Research Reagent Solutions for Biofouling Mitigation Studies
| Reagent/Material | Specification | Primary Function | Example Application |
|---|---|---|---|
| Carbon Fiber | AS4 7.1 μm diameter (Hexcel) [33] | Working electrode material for FSCV | Fabrication of CFMEs for adenosine detection |
| Silver Wire | 0.25 mm diameter (Sigma-Aldrich) [33] | Reference electrode base material | Construction of Ag/AgCl reference electrodes |
| Sodium Hypochlorite | 8.25% solution (Commercial bleach) [33] | Chloridizing agent for reference electrodes | Converting Ag wire to Ag/AgCl reference electrodes |
| Bovine Serum Albumin (BSA) | 40 g L⁻¹ in Tris buffer [32] | Biofouling simulation agent | In vitro testing of antifouling coatings |
| Nafion | 5% solution in solvent [33] | Cation-exchange polymer coating | Creating fouling-resistant barriers on electrodes |
| PEDOT:PSS | Conductive polymer dispersion [32] | Antifouling coating material | Electrodeposition on CFMEs for enhanced fouling resistance |
| Tris Buffer | 15 mM Tris HCl, 10 mM Tris base, pH 7.4 [33] [32] | Physiological buffer medium | Electrochemical measurements and fouling experiments |
| Natural Phosphate | Porous material for electrode modification [35] | Adenosine-selective adsorption | Modifying carbon paste electrodes for enhanced adenosine detection |
| Sodium Sulfide | 1 M stock in Tris buffer [32] | Reference electrode fouling agent | Testing reference electrode stability against sulfide exposure |
Successful mitigation of electrode biofouling and surface passivation in FSCV for adenosine detection requires a multifaceted approach. The strategies outlined herein—including three-electrode configuration, careful material selection, targeted surface coatings, and waveform optimization—collectively address the complex fouling mechanisms encountered in long-term implantation studies. Implementation of these protocols will significantly enhance the stability, sensitivity, and reliability of adenosine measurements, enabling more robust investigation of purinergic signaling in neurological function and dysfunction. Researchers should prioritize validation of these approaches in their specific experimental models to optimize parameters for unique application requirements.
Precise Ohmic Drop Compensation for Accurate Voltammogram Interpretation
In the field of electroanalytical chemistry, ohmic drop (or IR drop) represents a fundamental challenge that becomes critically pronounced during fast-scan cyclic voltammetry (FSCV). This phenomenon refers to the potential drop (i*R𝑢) that occurs when current (i) flows through the uncompensated solution resistance (R𝑢) between the working electrode surface and the reference point [38] [39]. In practical terms, the ohmic drop causes a discrepancy between the potential applied by the potentiostat and the actual potential experienced at the electrode-electrolyte interface [39]. For researchers investigating rapid adenosine signaling in neurological systems, this artifact can severely distort voltammograms, compromising the accuracy of both quantitative measurements and kinetic analyses [38] [11].
The significance of ohmic drop compensation becomes particularly evident in FSCV applications for adenosine detection, where scan rates of 400 V/s or higher are routinely employed to track subsecond neurochemical fluctuations [40] [11]. At these elevated scan rates, ohmic drop can reach tens or even hundreds of millivolts, potentially shifting peak potentials enough to obscure critical information about adenosine's redox behavior [38]. Without adequate compensation, the extracted thermodynamic and kinetic parameters may contain substantial errors, leading to flawed interpretations of adenosine release and clearance dynamics in brain regions such as the striatum and motor cortex [11].
Technical Challenges and Compensation Principles
Fundamental Challenges in Ohmic Drop Compensation
The pursuit of accurate ohmic drop compensation in FSCV confronts several technical hurdles that intensify with increasing scan rates. During fast scans (up to 1600 V/s or higher), the faradaic current intensifies proportionally to the square root of the scan rate for diffusion-controlled reactions, while the charging current increases linearly with scan rate [38] [40]. This relationship means that at the high scan rates essential for monitoring rapid adenosine transients, the ohmic drop distortion becomes magnified, potentially shifting redox peaks by several hundred millivolts [38]. In severe cases, these distortions can cause redox peaks to disappear entirely from the voltammetric window, rendering the electrochemical system uninterpretable [38].
Traditional positive feedback compensation techniques, which feed a portion of the output signal back to the potentiostat's input, represent the most common approach to addressing ohmic drop [38]. However, these methods suffer from a critical limitation: the compensation level is typically adjusted until the circuit reaches self-excited oscillation, which often occurs when the compensation exceeds 100% of the actual ohmic drop [38]. This overcompensation problem stems from the fact that the oscillation point depends not only on the solution resistance but also on circuit design, component layout, and the specific electrochemical system [38]. Consequently, even experienced researchers may achieve only approximate compensation, with errors of 5% or more resulting in significant potential shifts—approximately 50 mV for a 1 mA current flowing through 1 kΩ of uncompensated resistance [38].
Advanced Compensation Methodology
Recent technological advances have introduced more precise approaches to ohmic drop compensation, notably through direct online measurement of solution resistance prior to voltammetric analysis. This methodology employs dedicated instrumentation—typically incorporating impedance measurement chips such as the AD5933—to quantitatively determine the uncompensated solution resistance before initiating FSCV scans [38]. Following this measurement, the system automatically configures the positive feedback parameters to precisely match the measured resistance, enabling real-time iR𝑢 compensation during subsequent voltammetric experiments [38].
This two-step process represents a significant improvement over conventional compensation techniques because it decouples the resistance measurement from the compensation adjustment, eliminating the guesswork associated with traditional positive feedback approaches [38]. The precision offered by this method enables reliable FSCV at scan rates up to 1600 V/s in practical electrochemical systems, and even up to 2000 V/s in theoretical electrochemical cells [38]. For adenosine researchers, this level of compensation fidelity ensures that the subtle voltammetric features used to identify and quantify this neurochemical remain undistorted, even when detecting transient, spontaneous release events lasting mere seconds [11].
Table 1: Comparison of Ohmic Drop Compensation Techniques
| Compensation Method | Principle | Accuracy Limitations | Suitable Scan Rates |
|---|---|---|---|
| Traditional Positive Feedback [38] | Proportional feedback of output signal to potentiostat input | Self-excited oscillation occurs at >100% compensation; Highly system-dependent | Moderate scan rates (<100 V/s) |
| Current Interruption [38] | Measures potential decay during brief current cessation | Limited temporal resolution; Complex implementation | Lower scan rates |
| Online Resistance Measurement + Compensation [38] | Direct solution resistance measurement followed by precise feedback | Minimal error; Limited mainly by measurement precision | Very high scan rates (≥1600 V/s) |
Experimental Protocols for Ohmic Drop Compensation
Protocol 1: Online Solution Resistance Measurement and Compensation
This protocol describes the implementation of precise ohmic drop compensation through direct solution resistance measurement, adapted from methodology developed for ultrasensitive biosensing applications [38].
Materials and Equipment:
- Potentiostat system with digital control capabilities
- Solution resistance measurement module (e.g., based on AD5933 chip)
- Microcontroller unit (e.g., STM32F103ZET6)
- Digital potentiometer for feedback adjustment
- Standard three-electrode configuration: Working electrode (e.g., carbon-fiber microelectrode), Reference electrode (e.g., Ag/AgCl), Counter electrode
- Electrolyte solution appropriate for the system under study
Procedure:
- System Initialization: Power the microcontroller and potentiostat system. Initialize communication between all digital components.
- Resistance Measurement Phase:
- Activate the solution resistance measurement module prior to voltammetric scanning.
- Apply a small-amplitude AC signal across the working and counter electrodes using the integrated impedance chip.
- Measure the resulting current response and calculate the uncompensated solution resistance (R𝑢) through the microcontroller.
- Compensation Configuration:
- Translate the measured R𝑢 value into appropriate digital potentiometer settings using predefined calibration curves.
- Configure the positive feedback circuit with the calculated parameters to achieve precise iR𝑢 compensation.
- Voltammetric Analysis:
- Apply the selected waveform (e.g., triangular waveform from -0.4 V to +1.5 V and back at 400 V/s for adenosine detection) [11].
- Record the resulting current while maintaining the predetermined compensation settings.
- Validation:
- Verify compensation accuracy using a known redox standard (e.g., ferrocene) [38].
- Confirm minimal shift in peak separation (ΔEp) compared to theoretical values.
This protocol enables automated, precise ohmic drop compensation without requiring manual adjustment or achieving circuit oscillation, enabling accurate FSCV at scan rates up to 1600 V/s [38].
Protocol 2: Traditional Positive Feedback Compensation
For systems lacking dedicated resistance measurement capabilities, this protocol outlines standard positive feedback compensation procedures.
Materials and Equipment:
- Potentiostat with positive feedback compensation functionality
- Standard three-electrode configuration
- Electrolyte solution containing analyte of interest
Procedure:
- Initial Setup: Configure the potentiostat for FSCV with the desired waveform parameters.
- Baseline Acquisition:
- Record initial voltammograms without compensation to assess uncompensated ohmic drop.
- Note the peak separation (ΔEp) for reversible redox couples.
- Gradual Compensation:
- Systematically increase the positive feedback compensation level in small increments.
- After each adjustment, record voltammograms and observe the current response.
- Oscillation Point Detection:
- Continue increasing compensation until the circuit approaches self-excited oscillation.
- Reduce compensation slightly until oscillation ceases.
- Validation at Multiple Scan Rates:
- Perform voltammetry at different scan rates to verify consistent compensation.
- Compare ΔEp values to theoretical predictions for a reversible system (59.2/n mV at 25°C) [41].
Limitations Note: Researchers should be aware that this method typically results in overcompensation (≥105%), as oscillation generally occurs above the exact compensation point [38].
Figure 1: Workflow for precise ohmic drop compensation in FSCV experiments incorporating direct solution resistance measurement.
Research Reagent Solutions and Materials
Table 2: Essential Research Reagents and Materials for Ohmic Drop-Compensated FSCV
| Item | Function/Application | Example Specifications |
|---|---|---|
| Carbon-Fiber Microelectrodes [40] [11] | Working electrode for in vivo adenosine detection | 7µm diameter, cylindrical geometry, 65-75µm exposed length |
| Adenosine Standard Solutions [11] | Calibration and identification of adenosine signals | 10 mM stock in 0.1 M perchloric acid, diluted to 0.5-5 µM in Tris buffer |
| Supporting Electrolyte [11] | Provides ionic strength, minimizes unnecessary resistance | 15 mM Tris, 140 mM NaCl, 3.25 mM KCl, 1.2 mM CaCl₂, 1.25 mM NaH₂PO₄, 1.2 mM MgCl₂, 2.0 mM Na₂SO₄, pH 7.4 |
| Reference Electrode [11] [39] | Stable potential reference | Ag/AgCl electrode, prepared in 1 M HCl |
| Ferrocene Redox Standard [38] | Validation of ohmic drop compensation | 1-5 mM in appropriate non-aqueous solvent |
| Resistance Calibration Solutions [38] | Verification of resistance measurement accuracy | Known resistors (100-1000 Ω) or standard solutions |
Data Interpretation and Analysis
Recognizing Ohmic Drop Effects in Voltammetric Data
Proper interpretation of voltammograms requires careful attention to the characteristic signatures of insufficient ohmic drop compensation. The most evident indicator is an abnormally large peak separation (ΔEp) that exceeds the theoretical value of 59.2/n mV for a reversible system at 25°C [41]. This distortion becomes more pronounced with increasing scan rates and higher current magnitudes, providing an important diagnostic clue [41]. Additionally, researchers should suspect significant ohmic drop when observing asymmetric peak currents (ipa/ipc ≠ 1) without chemical complications, or when peak potentials shift systematically with changing scan rates in ways inconsistent with established electron transfer models [41].
For adenosine-specific FSCV employing waveforms from -0.4 V to +1.5 V at 400 V/s [11], uncompensated ohmic drop may manifest as variations in the characteristic oxidation peak around +1.3 V to +1.4 V, potentially leading to misidentification of spontaneous transients or inaccurate concentration estimates. Importantly, the effects of ohmic drop can be distinguished from slow electron transfer kinetics by examining analyte concentration dependence: ohmic drop distortions intensify with increasing concentration (and thus current), while kinetic effects remain concentration-independent [41].
Quantitative Assessment of Compensation Accuracy
Researchers should implement systematic validation procedures to quantify compensation accuracy following the application of ohmic drop correction methods. The most straightforward approach involves analyzing a well-characterized reversible redox system such as ferrocene under identical compensation settings [38]. The measured peak separation should approach the theoretical value of 59.2/n mV, with deviations of less than 5-10 mV indicating excellent compensation [41]. Additionally, the ratio of anodic to cathodic peak currents should approach unity (0.95-1.05) for a simple reversible system [41].
Table 3: Troubleshooting Guide for Ohmic Drop Compensation
| Observation | Potential Cause | Corrective Action |
|---|---|---|
| Increasing ΔEp with higher scan rates [41] | Incomplete ohmic drop compensation | Verify resistance measurement; Increase compensation level; Check electrode positioning |
| Distorted peak shapes [39] | Excessive positive feedback (overcompensation) | Slightly reduce compensation level until oscillation ceases |
| Unstable baseline [38] | Circuit approaching oscillation threshold | Adjust compensation slightly below oscillation point |
| Inconsistent results between electrodes [40] | Variations in electrode geometry or placement | Standardize electrode fabrication; Ensure consistent reference electrode positioning |
| Disappearing redox peaks [38] | Severe ohmic drop shifting peaks outside window | Increase compensation; Verify waveform limits encompass expected peak potentials |
Figure 2: Diagnostic and resolution pathway for addressing ohmic drop artifacts in FSCV experiments.
Application to Adenosine Detection Research
The implementation of precise ohmic drop compensation methodologies holds particular significance for FSCV studies investigating adenosine dynamics in neurological systems. The Venton lab and other research groups have demonstrated that adenosine operates on rapid timescales, with spontaneous release events occurring within seconds in brain regions such as the motor cortex and dorsal striatum [11] [15]. Detecting these transient fluctuations—which exhibit amplitudes of approximately 66-85 nM and frequencies of 0.5-1.5 Hz—demands exceptional voltammetric precision that can only be achieved with meticulous attention to uncompensated resistance effects [11].
For adenosine researchers employing the established triangular waveform from -0.4 V to +1.5 V at 400 V/s [11], even modest ohmic drop artifacts could significantly impact the identification and quantification of spontaneous transients. The characteristic adenosine voltammogram features distinct oxidation and reduction peaks that facilitate discrimination from other neurochemicals, but these identifying features become blurred without proper compensation [11] [15]. Furthermore, studies exploring adenosine release in response to external stimuli—such as the approximately 3-second tail pinch protocol—require uncompromised temporal resolution and quantitative accuracy to establish valid correlations between neurological events and purinergic signaling [11].
The development of digital circuits capable of precise ohmic drop compensation through online resistance measurement [38] represents a particularly valuable advancement for adenosine research, potentially enabling the detection of previously unobservable low-concentration transients and improving the reliability of concentration estimates in pathophysiological conditions such as stroke and seizure models [11] [15]. By implementing the compensation protocols outlined in this document, researchers can significantly enhance the fidelity of their adenosine measurements, contributing to a more accurate understanding of this crucial neuromodulator's rapid signaling mechanisms.
The accurate detection of neurotransmitters, such as adenosine, via Fast-Scan Cyclic Voltammetry (FSCV) is fundamentally linked to the physical and material characteristics of the implanted microelectrode. Conventional cylindrical carbon fiber microelectrodes (CFMEs) with diameters of 7 µm have been widely used for their high temporal resolution and biocompatibility. However, their application in chronic monitoring is limited by mechanical fragility and a propensity to cause tissue damage upon insertion, triggering neuroinflammatory responses that degrade signal quality over time [12] [42]. Recent research demonstrates that modifying electrode geometry, specifically through the implementation of cone-shaped designs, presents a robust strategy to enhance mechanical durability, improve biocompatibility, and extend functional longevity for adenosine and other neurochemical sensing [12] [9]. This application note details the protocol for fabricating cone-shaped microelectrodes and quantifies their performance advantages within the context of adenosine FSCV research.
Performance Data and Comparative Analysis
Recent experimental data directly compares the performance of traditional 7 µm cylindrical CFMEs, 30 µm cylindrical CFMEs, and novel 30 µm cone-shaped CFMEs. The quantitative results, summarized in the table below, highlight the significant benefits of the cone-shaped geometry.
Table 1: Comparative Performance of Cylindrical vs. Cone-Shaped Carbon Fiber Microelectrodes
| Performance Metric | 7 µm Bare CFME | 30 µm Bare CFME | 30 µm Cone-Shaped CFME |
|---|---|---|---|
| In Vitro Sensitivity (pA/µm²) | 12.2 ± 4.9 [12] | 33.3 ± 5.9 [12] | Data not explicitly stated; expected to be high due to larger base diameter. |
| In Vivo Dopamine Signal (nA) | 24.6 ± 8.5 [12] | 12.9 ± 8.1 [12] | 47.5 ± 19.8 [12] |
| Relative Lifespan (Erosion Test) | Baseline (1x) [12] | Data not explicitly stated. | 4.7-fold increase [12] |
| Glial Activation (Iba1/GFAP) | Moderate [12] | High [12] | Significantly lower [12] |
| Key Advantage | Minimal initial tissue damage [12] | High mechanical strength & in vitro sensitivity [12] | Superior biocompatibility, signal strength, and longevity [12] |
The data shows that while increasing the diameter of a cylindrical CFME to 30 µm improves mechanical robustness and in vitro sensitivity, it causes greater tissue damage in vivo, paradoxically reducing the detected neurotransmitter signal. The cone-shaped 30 µm CFME successfully mitigates this trade-off, combining the strength of a larger electrode with a geometry that minimizes insertion trauma, resulting in a 3.7-fold improvement in in vivo signal and a 4.7-fold increase in lifespan compared to standard 7 µm CFMEs [12].
Experimental Protocol: Fabrication of Cone-Shaped Carbon Fiber Microelectrodes
The following section provides a detailed methodology for fabricating 30 µm cone-shaped carbon fiber microelectrodes via electrochemical etching, a critical step for improving biocompatibility in adenosine FSCV studies.
Materials and Equipment
Table 2: Research Reagent Solutions for Cone-Shaped CFME Fabrication
| Item | Function/Description |
|---|---|
| 30 µm Carbon Fiber | Structural core of the microelectrode (Source: World Precision Instruments) [12]. |
| Tris Buffer (pH 7.4) | Electrolyte solution for the electrochemical etching process [12]. |
| Direct Current (DC) Power Supply | Provides the 10 V potential required for controlled electrolysis [12]. |
| Linear Actuator | Precisely controls the vertical withdrawal speed of the carbon fiber during etching to define the cone geometry [12]. |
| Microscalpel | For trimming the etched carbon fiber to the final exposed length of ~100 µm [12]. |
| FSCV Potentiostat | For electrode preconditioning and subsequent neurotransmitter detection (e.g., WaveNeuro series) [15]. |
Step-by-Step Procedure
- Electrode Mounting: Securely mount a 30 µm carbon fiber into a standard microelectrode holder, exposing approximately 1 mm of the fiber length.
- Etching Setup: Submerge the exposed carbon fiber tip into a bath of Tris buffer. Connect the DC power supply to apply a voltage between the carbon fiber (anode) and a counter electrode in the solution.
- Electrochemical Etching: Apply 10 V DC for 20 seconds [12]. Simultaneously, activate the linear actuator to withdraw the carbon fiber vertically from the electrolyte solution at a constant, controlled speed.
- Geometry Control: The combination of electrolysis and withdrawal causes the submerged portion of the fiber to erode and detach, while the meniscus region forms the cone. The actuator speed dictates the final cone height, which should be controlled to between 100 and 120 µm [12].
- Final Trimming: After etching, use a microscalpel to trim the cone-shaped tip to a final exposed length of ~100 µm [12].
- Preconditioning: Before use in FSCV, precondition the electrode by applying a triangular FSCV waveform (e.g., from –0.4 V to +1.5 V at 400 V/s, 30 Hz) to ensure a stable and active electrochemical surface [12].
The workflow for this fabrication process is outlined below.
Application in Adenosine Research
The cone-shaped microelectrode design is particularly advantageous for the detection of adenosine, a purine neuromodulator with a critical role in neuroprotection, sleep regulation, and seizure termination [4] [43]. FSCV allows for the direct detection of rapid, transient adenosine release with sub-second temporal resolution, which is essential for understanding its rapid modulatory roles [4] [15].
Adenosine exerts its effects primarily through A1 receptors, which are G protein-coupled receptors (GPCRs) that hyperpolarize neurons and inhibit the release of other neurotransmitters like dopamine and glutamate [7]. The following diagram illustrates this signaling pathway and its relevance to FSCV measurement.
The cone-shaped CFME enhances this research by providing a more stable and biocompatible interface. The reduced tissue damage and glial activation lead to a healthier local microenvironment, minimizing confounding signals from the inflammatory response and allowing for more accurate, long-term measurement of delicate physiological processes like rapid adenosine signaling [12] [44]. This design is therefore a promising tool for advancing studies of adenosine dynamics in both basic neuroscience and clinical contexts, such as monitoring adenosine release during deep brain stimulation or seizure activity [43].
Digital Filtering and Signal Processing to Isolate Adenosine from Background Drift and Noise
Within the framework of fast-scan cyclic voltammetry (FSCV) research for adenosine detection, a significant challenge is the reliable isolation of subtle adenosine signals from confounding background drift and electrical noise. FSCV provides sub-second temporal resolution for monitoring rapid adenosine dynamics in vivo, which is essential for understanding its neuromodulatory roles [4]. However, conventional data analysis techniques often fall short because they fail to utilize the full three-dimensional data structure of FSCV color plots and struggle with the evolving shape of adenosine's cyclic voltammogram during transient events [8]. This application note details a robust methodology combining digital filtering and image-based analysis to overcome these limitations, enabling precise, automated detection of adenosine with high specificity against common interferents.
The SSIM Image Analysis Framework for Adenosine Detection
Core Principle: Treating FSCV Data as Images
The Structural Similarity Index (SSIM) algorithm transforms FSCV data analysis by treating three-dimensional color plots (current vs. potential vs. time) as images [8]. Each neurotransmitter produces a unique "fingerprint" in these color plots based on its distinct redox properties. The SSIM index quantifies the similarity between a sample FSCV color plot and a reference adenosine color plot by comparing luminance, contrast, and structural patterns [8]. This approach leverages the entire dataset rather than focusing on limited current-time traces, resulting in significantly improved detection accuracy.
Adenosine is particularly challenging to detect because it undergoes multiple oxidation steps, leading to two Faradaic peaks that evolve on different timescales within the same transient event [8]. The SSIM method excels in detecting these shape-changing signals, outperforming traditional techniques like principal component regression (PCR) that analyze single cyclic voltammograms and struggle with dynamic CV shapes [8].
Performance Metrics and Advantages
The SSIM method demonstrates exceptional performance in automated adenosine detection, as quantified through rigorous validation:
Table 1: Performance Metrics of SSIM Analysis for Adenosine Detection
| Metric | Performance Value | Interpretation |
|---|---|---|
| Precision | 99.5 ± 0.6% | Extremely low false positive rate |
| Recall | 95 ± 3% | High detection rate of true events |
| F1 Score | 97 ± 2% | Excellent overall accuracy |
| Selectivity | Successfully rejects pH changes, histamine, and H2O2 | High specificity for adenosine |
This method has proven capable of detecting simultaneous adenosine and dopamine release events, demonstrating its utility for studying complex neurochemical interactions [8]. Furthermore, the implementation of digital filtering as a preprocessing step eliminates the need for background subtraction, simplifying the analysis workflow and enhancing signal integrity [8].
Experimental Protocols
Carbon-Fiber Microelectrode Preparation and FSCV Setup
Purpose: To fabricate and condition reliable sensors for adenosine detection.
- CFME Fabrication: Prepare carbon-fiber microelectrodes (CFMEs) from T-650 carbon fibers (7-μm diameter) sealed in glass capillaries with an exposed fiber length of 100 μm [8].
- Electrochemical Setup: Use a two-electrode system with a CFME working electrode and Ag/AgCl reference electrode connected to a potentiostat (e.g., ChemClamp) [8].
- FSCV Waveform Parameters:
- Holding potential: -0.4 V
- Switching potential: +1.45 V
- Scan rate: 400 V/s
- Repetition rate: 10 Hz [8]
- Electrode Conditioning: Precondition CFMEs before experiments using a 1.5 V FSCV sweep (-0.4 to 1.5 V at 400 V/s, 30 Hz) followed by application of the standard FSCV waveform [9].
Digital Filtering Protocol for Background and Noise Removal
Purpose: To eliminate background drift and high-frequency noise while preserving adenosine signals.
- High-Pass Filtering:
- Filter Type: Second-order Butterworth filter
- Cutoff Frequency: 0.03 Hz (for background detrending) [8]
- Implementation: Apply to the entire FSCV color plot to remove slow baseline drift without affecting transient adenosine signals.
- Noise Reduction Filtering:
SSIM Analysis Implementation
Purpose: To automate adenosine detection using image similarity analysis.
- Software Environment: Implement algorithms in MATLAB using the built-in SSIM function [8].
- Reference Selection:
- SSIM Calculation:
- Normalize both sample and reference color plots to their maximum current prior to comparison [8].
- Compute the SSIM index locally for each potential-time pixel to generate an SSIM matrix [8].
- Apply element-wise multiplication with a weight matrix emphasizing adenosine-specific features [8].
- Sum all elements in the product matrix and normalize to obtain the final SSIM index (ranging 0-1) [8].
- Threshold Optimization: Determine optimal SSIM cutoff scores through receiver operator characteristic (ROC) analysis to maximize true positives while minimizing false positives [8].
Visualization of Workflows
Adenosine Detection and Analysis Workflow
Signal Processing and Specificity Verification
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Research Reagents and Materials for FSCV Adenosine Detection
| Item | Specifications/Function | Source/Example |
|---|---|---|
| Carbon Fiber | T-650 fibers (7-μm diameter) for CFME fabrication [8] | Cytec Engineering Materials |
| Adenosine Standard | Preparation of stock solutions (10 mM in 0.1 M HClO4) [8] | Acros Organics |
| Buffer System | Phosphate-buffered saline (PBS): 131.25 mM NaCl, 3.00 mM KCl, 10.0 mM NaH2PO4, 1.2 mM MgCl2, 2.0 mM Na2SO4, 1.2 mM CaCl2 (pH 7.4) [8] | Standard chemical suppliers |
| Selectivity Controls | Histamine, H2O2, pH solutions (pH 7.3, 7.5) for interference testing [8] | Various chemical suppliers |
| Potentiostat | FSCV-capable instrument with two-electrode configuration | ChemClamp (Dagan) |
| Analysis Software | MATLAB 2019b with custom SSIM algorithm implementation [8] | MathWorks |
Discussion and Applications
The combination of digital filtering and SSIM image analysis represents a significant advancement for adenosine detection in FSCV research. This methodology enables researchers to study rapid adenosine signaling with unprecedented accuracy, which is crucial for understanding its neuromodulatory functions in processes such as neurotransmission, blood flow regulation, and sleep-wake cycles [4]. The technique's ability to detect spontaneous adenosine release events, which are transient and unpredictable, opens new avenues for investigating activity-dependent adenosine signaling in various neurological contexts.
Recent applications demonstrate the growing importance of this approach in neuroscience research. Multiplexing FSCV with genetically encoded fluorescent sensors has revealed simultaneous adenosine, dopamine, and glutamate interactions, showing adenosine's inhibitory effect on both neurotransmitters within a 250 μm radius via A1 receptors [7]. This spatial and temporal profiling of adenosine neuromodulation provides critical insights for developing targeted therapies for neurological disorders where adenosine signaling is disrupted.
The robustness of this methodology also supports its potential translation to clinical applications, including closed-loop deep brain stimulation systems that could use real-time adenosine monitoring as a feedback mechanism for optimizing therapeutic outcomes [9]. As FSCV technology continues to evolve with improved electrode designs and analytical approaches, the ability to reliably isolate adenosine signals from complex backgrounds will remain fundamental to advancing both basic neuroscience and therapeutic development.
Validating FSCV Signals and Comparing Analytical Techniques
The 'Five Golden Rules' for In Vivo Neurotransmitter Validation
The measurement of dynamic neurotransmitter signaling in the living brain is fundamental to understanding brain function in health and disease. Fast-scan cyclic voltammetry (FSCV) has emerged as a powerful technique for monitoring electroactive neurotransmitters with high temporal and spatial resolution, enabling researchers to detect neurotransmitter release and uptake events on a sub-second timescale [13] [45] [46]. Unlike other neurochemical techniques that suffer from poor temporal resolution, FSCV provides a unique window into the rapid dynamics of neurotransmission [47] [46].
When applied to the study of adenosine, a potent neuromodulator with roles in sleep regulation, neuroprotection, and seizure termination, FSCV has revealed a previously unappreciated rapid signaling mode that operates on a scale of seconds [4] [11]. However, the complex chemical environment of the brain presents significant challenges for specifically identifying neurotransmitter signals. To ensure scientific rigor and data validity, the neuroscience community established a set of validation criteria known as the "Five Golden Rules" for in vivo neurotransmitter identification [13] [46]. These rules provide a critical framework for confirming the chemical identity of signals detected via FSCV, particularly when translating these methods to clinical studies in humans [13].
This application note details the application of these five golden rules within the context of adenosine research using FSCV, providing detailed protocols and analytical approaches to ensure data accuracy and reproducibility.
The Five Golden Rules: Framework for Validation
The "Five Golden Rules" comprise a comprehensive validation strategy to confirm the chemical identity of neurotransmitters measured in vivo. These rules were developed to address the inherent challenges of selectivity in complex biological environments and remain foundational for rigorous electrochemical neuroscience research [46]. The following sections outline each rule with specific application to adenosine detection.
Rule 1: Electrochemical Signature
Principle: The first and most immediate line of identification involves confirming that the analyte produces a characteristic current profile at specific oxidation and reduction potentials when the voltage waveform is applied.
Application to Adenosine: Adenosine exhibits a distinct cyclic voltammogram characterized by a primary oxidation peak at approximately +1.5 V and a corresponding reduction peak near +1.0 V (vs. Ag/AgCl reference electrode) [11]. This signature is the primary feature used for initial identification during FSCV experiments.
Table 1: Characteristic Electrochemical Profiles of Common Neurotransmitters Detected via FSCV
| Neurotransmitter | Primary Oxidation Peak (V) | Reduction Peak (V) | Distinguishing Features |
|---|---|---|---|
| Adenosine | +1.5 | +1.0 | Broad oxidation profile [11] |
| Dopamine | +0.6 | -0.2 | Sharp, well-defined peaks [13] |
| Serotonin | +0.8 to +1.0 | N/A | Lack of prominent reduction peak [46] |
Experimental Protocol:
- Carbon Fiber Electrorode Preparation: Prepare a carbon-fiber microelectrode by aspirating a single carbon fiber (≈7 μm diameter) into a glass capillary. Use a pipette puller to create a sealed, tapered electrode. Trim the exposed fiber to 50-100 μm in length [11].
- In Vitro Calibration: Place the electrode in a flow injection system with Tris buffer (pH 7.4) at room temperature.
- FSCV Recording Parameters: Apply a triangular waveform from -0.4 V to +1.5 V and back, at a scan rate of 400 V/s, repeated every 100 ms [11].
- Background Subtraction: Record and subtract the background current to isolate the Faradaic current of the analyte.
- Signal Acquisition: Introduce a known concentration of adenosine (e.g., 1.0 μM) into the flow cell and record the resulting cyclic voltammogram. Compare this "fingerprint" to signals obtained in vivo.
Rule 2: Anatomical Validation
Principle: The brain region under investigation must have established neuroanatomy supporting the presence and release of the putative neurotransmitter.
Application to Adenosine: FSCV recordings of adenosine should be targeted to brain regions with documented adenosine signaling, such as the dorsal striatum and motor cortex, which express high densities of adenosine receptors (A1 and A2A) [11].
Experimental Protocol:
- Stereotaxic Surgery: Anesthetize the animal (e.g., with urethane, 1.3 g/kg i.p.) and secure it in a stereotaxic frame.
- Electrode Placement: Using established stereotaxic coordinates (e.g., from bregma: AP +0.4 mm, ML ±2.4 mm for striatum), lower the carbon-fiber electrode to the target depth (e.g., DV -4.5 to -5.0 mm for dorsal striatum) [11].
- Histological Verification: Upon experiment completion, perfuse the animal transcardially with formalin. Remove the brain, section it coronally, and stain with Cresyl Violet or a similar stain. Verify the recording site location microscopically. Only data from correctly placed electrodes should be considered valid.
Rule 3: Kinetic Validation
Principle: The temporal dynamics (release and uptake) of the measured signal must be consistent with the known kinetics of the neurotransmitter system under investigation.
Application to Adenosine: Spontaneous "transient" adenosine release events, as detected by FSCV, typically exhibit rapid onset and clearance, lasting from 1 to 5 seconds, with frequencies ranging from 0.5 to 1.5 Hz in anesthetized rats [11].
Experimental Protocol:
- Detection of Transients: Record spontaneous adenosine fluctuations for at least 30 minutes in the target brain region.
- Data Analysis: Use analysis software (e.g., Clampfit, GraphPad Prism, or custom Matlab scripts) to measure the amplitude, duration, and inter-event intervals of detected transients.
- Kinetic Parameters: Calculate the half-life and uptake rate constant for evoked or spontaneous adenosine release events to characterize the kinetic profile.
Rule 4: Pharmacological Validation
Principle: The measured signal should be manipulated in a predictable manner by drugs known to affect the synthesis, release, reception, or reuptake of the putative neurotransmitter.
Application to Adenosine: Pharmacological agents that block adenosine receptors or alter its metabolism should reliably alter the FSCV signal. For instance, application of an adenosine A1 receptor antagonist should modulate adenosine dynamics.
Table 2: Pharmacological Tools for Validating Adenosine Signals
| Drug/Tool | Target | Expected Effect on Adenosine Signal | Example Usage |
|---|---|---|---|
| DPCPX | A1 Receptor Antagonist | Alters concentration and dynamics via feedback mechanisms [11] | Local application via reverse microdialysis |
| NBTI | Adenosine Transporter Inhibitor | Slows clearance, increases amplitude and duration of signals [4] | Systemic or local administration |
| Enzyme Cocktails | Adenosine Deaminase | Degrades adenosine, eliminating detected signal | Addition to flow cell for in vitro verification |
Experimental Protocol:
- Baseline Recording: Establish a stable baseline of spontaneous or evoked adenosine transients for at least 30 minutes.
- Drug Administration: Administer the pharmacological agent systemically or locally via reverse microdialysis.
- Post-Drug Recording: Continue FSCV recordings for a sufficient duration to capture the drug's effect (e.g., 60-90 minutes).
- Data Comparison: Statistically compare key parameters (e.g., transient frequency, amplitude, half-life) before and after drug administration using paired t-tests or ANOVA.
Rule 5: Independent Validation
Principle: Whenever possible, the identity of the neurotransmitter should be confirmed using an independent, orthogonal technique.
Application to Adenosine: While challenging in vivo, especially in acute experiments, this can be approached by comparing FSCV signals with those from techniques like microdialysis coupled to HPLC, albeit with the recognition of microdialysis's lower temporal resolution [47] [46].
Experimental Protocol:
- Parallel Measurement: In a separate cohort of animals, implant a microdialysis probe in the same brain region targeted for FSCV.
- Sample Collection: Collect dialysate samples at a flow rate of 2 μL/min, typically over 5-10 minute intervals.
- HPLC Analysis: Analyze dialysate samples using HPLC with UV or mass spectrometry detection to quantify absolute adenosine concentrations.
- Data Correlation: While direct temporal correlation is difficult, the confirmed presence of adenosine in the region via microdialysis provides independent anatomical and chemical validation for the signals detected by FSCV.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials and Reagents for FSCV-based Adenosine Detection
| Item | Function/Description | Example/Specification |
|---|---|---|
| Carbon Fiber Microelectrode | Working electrode for neurotransmitter detection; 7 μm diameter carbon fiber provides high spatial resolution and sensitivity [11]. | T-650 Carbon Fiber (Cypress) in glass capillary, 50-100 μm exposed tip length |
| Ag/AgCl Reference Electrode | Provides a stable reference potential for the electrochemical cell, critical for accurate voltage application [11]. | Chloridized silver wire, implanted in contralateral brain hemisphere |
| Potentiostat | Instrument that applies the voltage waveform and measures the resulting current. | UNC Vector Voltammeter or equivalent; capable of high-speed scans (400 V/s) |
| Triangular Waveform | Voltage profile used to oxidize and reduce the analyte, generating the characteristic voltammogram [11]. | -0.4 V to +1.5 V vs. Ag/AgCl, 400 V/s scan rate, repeated at 10 Hz |
| Data Acquisition System | Hardware and software for digitizing, visualizing, and storing current signals. | National Instruments DAQ card with LabVIEW software (e.g., Tar Heel CV) |
| Calibration Solutions | Used for post-hoc calibration of electrode sensitivity to adenosine. | 0.5 - 5 μM adenosine in Tris buffer, pH 7.4 [11] |
Experimental Workflow and Signaling Pathways
The following diagrams outline the core experimental workflow for FSCV and the adenosine signaling pathway, integrating the validation rules into the research process.
Figure 1: Experimental Workflow for FSCV Adenosine Validation
Figure 2: Adenosine Signaling and Detection Pathway
The "Five Golden Rules" provide an indispensable, systematic framework for validating neurotransmitter identity in FSCV experiments. Their rigorous application is paramount for generating reliable and reproducible data, particularly in the context of adenosine research where rapid signaling dynamics are now known to play critical functional roles [4] [11]. As FSCV continues to evolve and finds new applications in clinical settings [13], adherence to these foundational principles will ensure scientific rigor and the accurate interpretation of complex neurochemical events in the brain.
The study of neuromodulators such as adenosine is crucial for understanding brain function and developing treatments for neurological disorders. The selection of an appropriate in vivo monitoring technique directly influences the validity and interpretation of experimental data. Fast-scan cyclic voltammetry (FSCV) and microdialysis represent two principal approaches with complementary strengths and limitations. This application note provides a detailed comparison of these techniques, focusing on their temporal and spatial resolution characteristics, within the broader context of adenosine detection research. The content is structured to assist researchers, scientists, and drug development professionals in selecting and implementing the optimal methodology for their specific investigative needs.
Technical Comparison: FSCV vs. Microdialysis
The core specifications of FSCV and microdialysis differ significantly, making each technique suitable for distinct experimental questions. The following table summarizes their key performance characteristics based on current literature.
Table 1: Key Performance Characteristics of FSCV and Microdialysis
| Characteristic | Fast-Scan Cyclic Voltammetry (FSCV) | Microdialysis |
|---|---|---|
| Temporal Resolution | Sub-second to seconds (100 ms scan rate) [20] [48] | Minutes to tens of minutes (5-30 min typical sample collection) [49] [48] |
| Spatial Resolution | Micron-scale (Carbon fiber electrode: ~7 µm diameter) [50] | Millimeter-scale (Probe diameter: ~200-300 µm) [50] |
| Principal Measurement | Transient, rapid concentration changes [48] | Time-averaged, basal extracellular concentration [51] |
| In Vivo Selectivity | Based on electrochemical signature (e.g., oxidation peaks); can distinguish adenosine via its two characteristic oxidation peaks [20] | Based on physical separation; can be combined with HPLC/LC-MS for high specificity [52] |
| Key Advantage | Excellent for monitoring spontaneous, transient release events (e.g., adenosine transients) with high spatiotemporal precision [20] [53] | Broad chemical scope; capable of monitoring multiple purines (e.g., adenosine, inosine, hypoxanthine) simultaneously when coupled with separation techniques [49] |
| Primary Limitation | Limited to electroactive analytes; complex data analysis for random transients [50] [20] | Poor temporal resolution; significant tissue damage and foreign body response that can alter basal analyte levels [50] [51] |
| Typical LOD for Adenosine | Demonstrated for transient release events in vivo [20] [53] | Low nanomolar range (e.g., 0.02 nM LLOQ reported in LC-MS/MS methods) [52] |
Experimental Protocols
Protocol for Detecting Adenosine Transients Using FSCV
This protocol details the use of FSCV for the detection of spontaneous adenosine release, which produces transient events lasting only a few seconds [20].
1. Electrode Preparation and Calibration:
- Fabricate a carbon-fiber microelectrode by sealing a single carbon fiber (e.g., 7 µm diameter) in a glass capillary [50] [48].
- Prior to implantation, calibrate the electrode in vitro using a flow injection system with a known concentration of adenosine (e.g., 1 µM) in artificial cerebrospinal fluid (aCSF).
- For adenosine-specific detection, apply a triangular waveform. A typical waveform ramps from a holding potential of -0.4 V to a switching potential of 1.5 V and back, at a scan rate of 400 V/s, repeated every 100 ms [20] [53].
- Identify the voltages for the primary (∼1.4 V) and secondary (∼1.0 V) oxidation peaks of adenosine from the background-subtracted cyclic voltammogram (CV). These voltages are used for subsequent analysis [20].
2. In Vivo Implantation and Data Acquisition:
- Anesthetize the animal (e.g., rat or swine) or use a freely moving preparation based on the experimental design.
- Stereotactically implant the calibrated carbon-fiber microelectrode into the brain region of interest (e.g., cortex or striatum).
- Implant a reference electrode (e.g., Ag/AgCl) at a remote site.
- Secure the electrode and begin continuous FSCV data acquisition using a suitable system (e.g., Wireless Instantaneous Neurotransmitter Concentration Sensor - WINCS) [53].
3. Data Analysis with Automated Algorithm:
- Perform incremental background subtraction on the raw data to reveal Faradaic current changes [20].
- Use an automated algorithm to identify adenosine transients from the current vs. time traces at the predetermined primary and secondary peak voltages. The algorithm applies the following rules [20]:
- A peak must be present at both the primary and secondary oxidation voltages.
- The secondary peak must lag the primary peak by 0.1 - 2.5 s.
- The ratio of the secondary to primary peak currents must fall between 0.49 and 0.89.
- The duration of the secondary peak must be longer than that of the primary peak.
- The signal-to-noise ratio (S/N) for both peaks must be greater than 3.
- The algorithm outputs the transient event time, concentration (from calibration of the primary peak), and duration.
Protocol for Measuring Basal Adenosine Using Microdialysis
This protocol describes the use of microdialysis to measure basal levels of adenosine in the brain extracellular fluid, which are typically in the low nanomolar range [52].
1. Probe Preparation and Implantation:
- Select a microdialysis probe with a suitable membrane molecular weight cutoff (e.g., 20 kDa) and membrane length (e.g., 1-4 mm). Note that probe membrane surface area is a variable that can affect recovery [51].
- Perfuse the probe with an isotonic perfusion fluid (e.g., aCSF) at a slow, constant flow rate (e.g., 1-2 µL/min) using a high-precision syringe pump. Note: The flow rate significantly influences measured adenosine concentration, with higher flow rates yielding lower concentrations [51].
- Anesthetize the animal and stereotactically implant the guide cannula. Insert the microdialysis probe into the target brain region through the cannula. Allow a post-surgical recovery period (often 24 hours) for the stabilization of basal analyte levels, as the implantation causes tissue trauma and a foreign body response that perturbs the local neurochemical environment [50] [52].
2. Sample Collection:
- Following the recovery period, begin collection of dialysate samples. Collect samples into vials containing a small volume of preservative (e.g., antioxidant or chelating agent) if necessary, to prevent analyte degradation.
- The collection interval determines temporal resolution. For basal measurements, intervals of 10-30 minutes are common, yielding sample volumes of 10-60 µL at a 1-2 µL/min flow rate [51] [49].
3. Sample Analysis via LC-MS/MS:
- This protocol uses Liquid Chromatography with Tandem Mass Spectrometry (LC-MS/MS) for high sensitivity and specificity [52].
- Sample Preparation: Mix a volume of the dialysate sample (e.g., 20 µL) with an internal standard (e.g., 5 µL of 50 nM 2-chloroadenosine) [52].
- Chromatographic Separation: Inject the sample onto a reverse-phase LC column (e.g., C18). Use a gradient elution with a mobile phase consisting of water and acetonitrile, both containing a volatile buffer such as ammonium acetate.
- Mass Spectrometric Detection: Use an electrospray ionization (ESI) source in positive ion mode. Monitor the specific transition for adenosine: precursor ion m/z 268 → product ion m/z 136. For the internal standard 2-chloroadenosine, monitor m/z 302 → m/z 170 [52].
- Quantification: Calculate the concentration of adenosine in the dialysate by comparing the peak area ratio of adenosine to the internal standard against a linear calibration curve.
4. Data Consideration:
- Reported basal adenosine concentrations in dialysates vary widely (0.8 - 2100 nM), influenced by factors such as flow rate and the use of anesthesia [51]. Anaesthesia has been shown to increase measured adenosine concentrations compared to freely behaving animals [51].
The Scientist's Toolkit: Essential Research Reagents and Materials
Successful implementation of FSCV or microdialysis requires specific reagents and instrumentation. The following table lists key solutions and materials used in the featured experiments.
Table 2: Key Research Reagent Solutions and Materials
| Item Name | Function / Application | Specifications / Notes |
|---|---|---|
| Carbon Fiber Microelectrode | FSCV working electrode for in vivo detection. | ~7 µm diameter, 100-400 µm active length [50]. Provides high spatial resolution and minimal tissue damage. |
| Triangular Waveform (FSCV) | Electrochemical stimulation for adenosine-selective detection. | E.g., -0.4 V to +1.5 V, 400 V/s, 10 ms duration, 100 ms interval [20] [53]. Generates adenosine's characteristic two-peak CV. |
| Adenosine Automated Algorithm | Software for identifying spontaneous adenosine transients in FSCV data. | Reduces analysis time from ~18 hours to 40 minutes per experiment. Filters based on peak lag, current ratio, and duration [20]. |
| Microdialysis Probe | In vivo sampling of the extracellular space. | ~300 µm diameter, 1-4 mm membrane length [50] [51]. Causes initial tissue penetration injury. |
| Artificial Cerebrospinal Fluid (aCSF) | Perfusion fluid for microdialysis; physiological buffer. | Isotonic solution, pH ~7.4. Carriers collected analytes to the sample vial [51]. |
| LC-MS/MS System | Off-line analysis of microdialysates for adenosine. | Provides high sensitivity (LLOQ ~0.05 nM) and specificity [52]. Uses MRM for quantitation (m/z 268→136). |
| 2-Chloroadenosine | Internal Standard (IS) for LC-MS/MS quantitation. | Added to dialysate samples to correct for variability in sample processing and instrument response [52]. |
The choice between FSCV and microdialysis is dictated by the specific research question. FSCV is unparalleled for investigating the dynamics of rapid, transient adenosine signaling, such as spontaneous release events or the precise correlation of adenosine surges with seizure termination, where its sub-second temporal resolution is critical [20] [53]. In contrast, microdialysis is the preferred method for quantifying stable, basal levels of adenosine over longer periods and for profiling multiple metabolites simultaneously, despite its poorer temporal resolution and more invasive nature [51] [49] [52].
In summary, FSCV offers high temporal and spatial resolution for monitoring rapid adenosine transients, whereas microdialysis provides a broader chemical profile of the extracellular environment at the cost of speed and spatial precision. Advances in sensor technology, such as the development of automated detection algorithms for FSCV [20], and the integration of microdialysis with faster analytical techniques like microchip electrophoresis [49], continue to push the boundaries of in vivo neurochemical monitoring. A clear understanding of the capabilities and limitations of each technique ensures the generation of robust and physiologically relevant data in adenosine research and drug development.
The precise detection of adenosine, a purine nucleoside with critical neuromodulatory and neuroprotective functions, is essential for understanding its role in both normal brain physiology and pathological states. No single analytical technique provides a complete picture of adenosine's complex spatiotemporal dynamics. Fast-scan cyclic voltammetry (FSCV) offers excellent temporal resolution for monitoring rapid adenosine fluctuations but provides limited chemical scope. Microdialysis enables broad neurochemical profiling yet suffers from poor temporal resolution. Genetically encoded sensors facilitate cell-specific monitoring with high spatial precision but require genetic modification. This protocol details a cross-platform validation framework that integrates these complementary methodologies to overcome their individual limitations, providing researchers with a comprehensive approach for verifying adenosine measurements across multiple detection platforms within the context of a broader thesis on FSCV for adenosine detection research.
Comparative Analytical Profiles
The integration of FSCV, microdialysis, and genetically encoded sensors creates a powerful analytical system where each component validates and complements the others. The table below summarizes the core characteristics of each technique.
Table 1: Technical comparison of adenosine detection methodologies
| Methodology | Temporal Resolution | Spatial Resolution | Key Advantages | Primary Limitations |
|---|---|---|---|---|
| FSCV | Sub-second [4] | ~10 µm (limited by microelectrode size) [50] | • Excellent temporal resolution• Direct, real-time measurement• Established in vivo protocols [15] [13] | • Limited chemical scope (electroactive analytes only)• Electrode fouling• Tissue damage from insertion [12] [13] |
| Microdialysis | Minutes | ~200-300 µm (probe diameter) [50] | • Broad neurochemical profile• No requirement for electroactive analytes• Sample collection for downstream analysis [50] | • Poor temporal resolution• Significant tissue damage and foreign body response• Low spatial resolution [50] |
| Genetically Encoded Sensors (e.g., HypnoS) | Sub-second to seconds [54] | Cellular and subcellular | • Cell-type specific targeting• High spatial resolution• Minimal physiological disruption when expressed• Intracellular measurement capability [54] | • Requires genetic modification• Limited to engineered model systems• Photobleaching potential• Calibration in vivo challenges [54] |
Integrated Experimental Protocols
Animal Preparation and Surgical Considerations
Conduct all procedures in accordance with institutional animal care guidelines. For simultaneous triple-platform recordings, anesthetize adult Sprague-Dawley rats (250-350 g) using urethane (1.5 g/kg i.p.) or isoflurane (1-3% in O₂). Secure the animal in a stereotaxic frame, maintain body temperature at 37°C using a heating pad, and confirm anesthetic depth throughout by monitoring pedal reflexes. Perform a midline scalp incision and drill craniotomies appropriate for your target brain region (e.g., striatum: AP +1.2 mm, ML ±2.0 mm from bregma). The integration of multiple probes necessitates careful planning of insertion coordinates and trajectories to minimize collective tissue damage while ensuring spatial overlap of measurement zones.
FSCV for Adenosine Detection
Carbon Fiber Microelectrode (CFME) Preparation
- Carbon Fiber Implementation: Utilize either conventional 7 µm diameter carbon fibers (AS4, Hexcel) for minimal tissue displacement or 30 µm diameter fibers (World Precision Instruments) with cone-shaped etching for enhanced mechanical durability [12]. The cone-shaped modification is achieved via electrochemical etching in Tris buffer at 10 V for 20 seconds while gradually retracting the fiber, creating a tip that reduces insertion-induced tissue damage [12].
- Electrode Fabrication: Seal a single carbon fiber into a silica tubing using epoxy resin, then trim the exposed fiber to 100-150 µm length using a surgical blade [12].
- Electrochemical Preconditioning: Before implantation, precondition CFMEs using FSCV waveforms (−0.4 V to +1.5 V, 400 V/s, 30 Hz) applied for 15-20 minutes in sterile PBS, followed by transition to standard adenosine detection waveform (−0.4 V to +1.45 V, 400 V/s, 10 Hz) until stable background current is achieved [15].
Data Acquisition and Analysis
- FSCV Parameters: Employ the following waveform parameters for adenosine detection: scanning from −0.4 V to +1.45 V and back at 400 V/s, applied at 10 Hz frequency. These parameters optimize the oxidation current for adenosine while maintaining electrode stability [15].
- Signal Processing: Collect currents using a high-gain amplifier (e.g., 400 nA/V). Apply background subtraction by using the average of 50 cyclic voltammograms collected before adenosine release as reference. Identify adenosine via its characteristic oxidation peak at approximately +1.25 V and reduction peak near −0.1 V versus Ag/AgCl reference [15].
- Calibration: Post-experiment, calibrate electrodes in solutions containing known adenosine concentrations (0.5-20 µM) in artificial cerebrospinal fluid (aCSF). Calculate sensitivity as current (nA) per µM adenosine.
Microdialysis Integration
Probe Implantation and Operation
- Probe Selection: Use concentric microdialysis probes with 3-4 mm membrane length and 220 µm outer diameter to minimize tissue disruption while maintaining adequate recovery [50].
- Spatial Configuration: Position the microdialysis probe approximately 500 µm from the FSCV electrode to ensure sampling from adjacent tissue volumes while preventing physical interference [50].
- Perfusion Parameters: Perfuse with aCSF (pH 7.4) containing (in mM): 145 NaCl, 2.7 KCl, 1.2 CaCl₂, 1.0 MgCl₂, 0.5 NaH₂PO₄ at 1.0 µL/min. After 60-90 minute equilibration period, collect dialysate samples every 5-10 minutes into microvials containing 5 µL of 0.1 N HCl to prevent adenosine degradation [50].
Analytical Validation
- HPLC Analysis: Separate adenosine using a C18 reverse-phase column (2.1 × 100 mm, 1.8 µm) with mobile phase of 50 mM ammonium acetate (pH 5.5) and 3% methanol at 0.2 mL/min.
- Detection: Quantify adenosine via UV detection at 260 nm or electrochemical detection at +1.4 V. Compare retention times with adenosine standards (1-500 nM) for quantification.
- Recovery Determination: Calculate relative recovery by retrodialysis: perfuse with 100 nM adenosine for 30 minutes pre-experiment and compare inlet-outlet concentrations. Typical recovery rates range from 10-20% for small molecules [50].
Genetically Encoded Sensor Implementation
Sensor Expression
- Sensor Selection: Utilize the recently developed HypnoS (Hypersensitive intracellular adenosine Sensor) for intracellular adenosine monitoring, which exhibits high sensitivity (EC₅₀ = 10.9 µM), excellent specificity for adenosine over related nucleotides, and sub-second response kinetics [54].
- Viral Delivery: For in vivo applications, administer AAV vectors carrying HypnoS under cell-specific promoters (e.g., CaMKIIα for neurons, GFAP for astrocytes) via stereotaxic injection (1-2 µL at 0.2 µL/min) 3-4 weeks prior to experiments to allow adequate expression [54].
- Control Experiments: Express the Ado-insensitive control HypnoS-mut (V200G and G201K mutations) in separate animals to distinguish specific adenosine responses from potential motion artifacts or nonspecific fluorescence changes [54].
Imaging and Quantification
- Optical Setup: Perform imaging using a two-photon microscope with a Ti:Sapphire laser tuned to 930 nm for optimal HypnoS excitation. Collect emission through a 525/50 nm bandpass filter [54].
- Data Acquisition: Acquire time-series images at 2-4 Hz frame rate. For ratiometric quantification, alternate excitation between 800 nm and 930 nm when using two-photon microscopy to enable pH-independent adenosine measurements [54].
- Calibration: Construct in vivo calibration curves by applying adenosine kinase inhibitor 5-Iodotubercidin (5-ITu, 5 µM) to elevate endogenous adenosine levels, or by iontophoretic adenosine application (100 nA, 5-10 s) while recording fluorescence changes [54].
Integrated Workflow and Signaling Pathways
The complementary nature of these techniques creates a powerful validation framework. The following diagram illustrates the experimental workflow and how signals from each platform interact:
This integrated approach reveals adenosine signaling pathways with unprecedented resolution. The following diagram maps the cellular pathways of adenosine production, release, and detection that can be investigated through this multi-platform approach:
The Scientist's Toolkit: Essential Research Reagents
Successful implementation of this cross-platform approach requires specific reagents and materials. The following table details the essential components:
Table 2: Key research reagents for adenosine detection methodologies
| Category | Specific Reagent/Item | Function/Application | Key Characteristics |
|---|---|---|---|
| FSCV Materials | Carbon fiber (AS4, 7 µm) [12] | Working electrode core | Small diameter minimizes tissue damage; suitable for acute studies |
| Carbon fiber (30 µm, cone-shaped) [12] | Working electrode for chronic studies | Enhanced mechanical durability; 4.7-fold increase in lifespan; reduced fouling | |
| Tris buffer (pH 7.4) [12] | Electrochemical medium | Electrochemical stability for in vitro calibration | |
| Microdialysis Supplies | Concentric microdialysis probes [50] | In vivo sampling | 220 µm diameter; 3-4 mm membrane length; minimal tissue disruption |
| Artificial cerebrospinal fluid [50] | Perfusion fluid | Physiological ion composition (Na⁺, K⁺, Ca²⁺, Mg²⁺, Cl⁻) | |
| 5-Iodotubercidin (5-ITu) [54] | Adenosine kinase inhibitor | Elevates endogenous adenosine for calibration (5 µM) | |
| Genetic Sensor Tools | HypnoS sensor plasmid [54] | Intracellular adenosine monitoring | EC₅₀ = 10.9 µM; 929% dynamic range; sub-second kinetics |
| AAV vectors (serotype 9) [54] | In vivo gene delivery | High neuronal transduction efficiency; minimal immunogenicity | |
| HypnoS-mut control [54] | Specificity control | Adenosine-insensitive variant; identifies nonspecific responses | |
| Analytical Standards | Adenosine (≥99% purity) | Calibration standard | Primary reference material for all platforms |
| HPLC mobile phase [50] | Chromatographic separation | 50 mM ammonium acetate (pH 5.5) with 3% methanol |
Data Integration and Validation Framework
Temporal Alignment and Correlation
The primary challenge in cross-platform validation lies in reconciling the different temporal resolutions of each technique. Implement the following alignment strategy:
- Time Normalization: Convert all data streams to a common timebase (seconds). Apply appropriate smoothing algorithms to higher temporal resolution data (FSCV, sensor) when comparing with microdialysis data.
- Event Markers: Introduce precisely timed physiological or electrical stimuli (e.g., 1-second forepaw stimulation, 10-second hypoxia exposure) to create alignment points across all recording modalities.
- Lag Compensation: Account for the inherent time lag (1-2 minutes) in microdialysis measurements due to tubing dead volume and membrane transit time when correlating with FSCV and sensor data.
Quantitative Cross-Validation
- Concentration Verification: Compare absolute adenosine concentrations obtained via microdialysis (after probe recovery correction) with FSCV measurements (post-calibration) during stable periods. Expect a strong correlation (R² > 0.85) when proper calibration procedures are followed.
- Dynamic Response Validation: Analyze response timing and amplitude across platforms following evoked release events. FSCV should detect adenosine transients within seconds, while genetically encoded sensors show intracellular correlates, and microdialysis captures the integrated concentration change over minutes.
- Cell-Type Specific Contribution: Utilize cell-specific promoter expression of genetic sensors to attribute adenosine dynamics to particular cellular sources (neurons vs. astrocytes), then correlate these with FSCV measurements of extracellular adenosine to infer cellular origins of release events [54].
Technical Considerations and Limitations
- Spatial Resolution Mismatch: Acknowledge that each technique samples different tissue volumes. FSCV measures within microns of the electrode surface, microdialysis samples a larger sphere around the probe, and genetic sensors report from individual cells.
- Interference Management: FSCV is susceptible to interference from pH shifts, oxygen changes, and other electroactive species. Use pharmacological validation (receptor antagonism) and background subtraction techniques to confirm adenosine identity [13].
- Tissue Impact Assessment: Recognize that both FSCV and microdialysis probes create some degree of tissue disruption. Utilize cone-shaped electrode geometries to minimize insertion damage and allow adequate recovery time (60-90 minutes) post-implantation before data collection [12] [50].
This integrated validation framework provides researchers with a robust methodology for confirming adenosine measurements across multiple detection platforms, significantly strengthening conclusions drawn from any single technique. The complementary nature of these approaches enables a more comprehensive understanding of adenosine dynamics than could be achieved by any method alone.
The translation of preclinical research on adenosine, a ubiquitous purine nucleoside with significant neuromodulatory and vascular functions, into safe and effective human therapies presents a formidable scientific challenge. Animal models have long been indispensable in biomedical research, yet significant physiological differences between rodents and humans often create a "translational gap" that hinders clinical application. Large animal models, particularly swine, have emerged as a crucial intermediary in the pathway from basic science to clinical implementation, especially for complex physiological measurements and technique validation [55] [43]. Their anatomical and physiological similarities to humans, including comparable body size, organ dimensions, metabolic processes, and pathophysiology, make them exceptionally well-suited for refining biomedical procedures and medical equipment [55]. This application note delineates the structured pathway from validation in large animal models to human feasibility studies for adenosine detection using fast-scan cyclic voltammetry (FSCV), providing researchers with detailed protocols and methodological considerations for successful translational research.
The attractiveness of porcine models in adenosine research extends beyond mere anatomical similarities. Pigs are monogastric omnivores that can develop obesity and dyslipoproteinemia similar to humans, and they exhibit comparable pharmacokinetics for orally or subcutaneously administered compounds [55]. Furthermore, pigs mature relatively quickly (6-7 months), have a short gestation period (approximately 114 days), high fecundity (8-14 offspring per litter), and a long life cycle, making production of genetically modified lines less time-consuming compared to other large animal species [55]. These characteristics, combined with wider public acceptance for their humane use in research compared to non-human primates, position swine as an optimal transitional model for adenosine research aimed at clinical application.
Technical Foundations: Fast-Scan Cyclic Voltammetry for Adenosine Detection
Fundamental Principles of FSCV
Fast-scan cyclic voltammetry (FSCV) is an electrochemical technique that has been refined over three decades for the detection of electroactive neurotransmitters and neuromodulators in biological systems. The method operates by applying a rapid, cyclic triangular waveform of electric potential between a working electrode (typically a carbon fiber microelectrode) and a reference electrode, resulting in the oxidation and/or reduction of electroactive analytes at characteristic potentials [43]. The resulting electrical currents have amplitudes proportional to the concentration of the analyte in the extracellular space, enabling estimation of neurotransmitter levels with high temporal resolution (sub-second), spatial precision (micrometer scale), sensitivity, and chemical selectivity [43]. For adenosine detection specifically, FSCV has revealed a previously unappreciated rapid signaling mode lasting only seconds, contrasting with the traditional understanding of adenosine as a tonic modulator operating on minute-to-hour timescales [4].
The application of FSCV in clinical environments requires meticulous attention to technical barriers that must be addressed to ensure both scientific rigor and patient safety. As noted in recent literature, "clinical questions that can be currently answered with FSCV are limited by technical barriers that must be addressed to ensure that clinical studies have appropriate scientific rigor and mitigate risks to patient safety" [43]. These challenges include potential confounding factors from interferant molecules with similar oxidation/reduction potentials, electrode biofouling, shifts in pH and ionic concentrations, increased oxygenated blood flow in the electrode microenvironment, and electrical or motion artifacts that can create signals mimicking neurotransmitter release [43].
Table 1: Key FSCV Parameters for Adenosine Detection in Animal Models
| Parameter | Typical Setting for Adenosine | Variations Reported | Functional Impact |
|---|---|---|---|
| Waveform Type | Triangular | Triphasic triangular (Earl et al.) | Optimizes detection specificity |
| Voltage Range | -0.4 V to +1.5 V | -0.2 V to +0.6 V (Cheney-Thamm et al.) | Affects oxidation/reduction profile |
| Scan Rate | 400 V/s | 10-900 V/s across studies | Influences temporal resolution and sensitivity |
| Scan Frequency | 10 Hz | 2 Hz (Earl et al.) | Balances temporal resolution with electrode stability |
| Resting Potential | -0.4 V | 0 V (Earl et al.) | Affects adsorption characteristics |
FSCV Validation Framework: The "Five Golden Rules"
To ensure reliability of in vivo neurotransmitter measurements, the FSCV community has established a set of validation guidelines known as the "Five Golden Rules" [43]:
- Identification of neurotransmitter-specific electrochemical signatures through analysis of cyclic voltammograms.
- Additional confirmation of chemical identity (e.g., through microdialysis at the FSCV recording site).
- Anatomical validation of the recording location via histological verification.
- Kinetic validation of spontaneous or evoked changes in neurotransmitter concentration.
- Pharmacological validation using receptor agonists/antagonists or uptake inhibitors.
These validation principles remain equally crucial when transitioning from small to large animal models and eventually to human studies, though their implementation may require adaptation to clinical constraints.
Figure 1: Pathway from Preclinical Development to Clinical Application for FSCV-based Adenosine Detection
Large Animal Models: Bridging the Translational Gap
Advantages of Porcine Models in Adenosine Research
The limitations of rodent models in predicting human physiological responses have been extensively documented. In a systematic review, Henderson et al. identified significant challenges in translating preclinical findings to human applications, with numerous drugs passing preclinical trials but failing in human testing [56]. Porcine models effectively address many of these limitations through several key advantages:
- Physiological Similarity: Swine share relevant metabolic physiology and pathophysiology with humans, including similar pancreatic and islet architecture crucial for metabolic studies [55].
- Technical Compatibility: Their comparable body size and anatomy enable the use of identical surgical procedures and medical devices intended for human application [55].
- Genetic Manipulability: The availability of genetic engineering tools facilitates the development of transgenic porcine models that better recapitulate human disease states [55].
These advantages are particularly relevant for adenosine research, given adenosine's dual roles in neurological and cardiovascular function across species. The successful use of swine in developing the bi-hormonal bionic pancreas technology demonstrates their value in complex physiological research [55]. This technology was initially established using a large animal diabetic porcine model before progressing to clinical trials [57] [58].
Table 2: Large Animal Models in FSCV Research for Adenosine Detection
| Species | Study Focus | FSCV Parameters | Key Findings | Translational Relevance |
|---|---|---|---|---|
| Swine (Sus scrofa domesticus) | STN-evoked striatal dopamine release [43] | Triangular waveform -0.4 to +1.5 V, 400 V/s, 10 Hz | STN electrical stimulation evoked intensity and frequency dependent striatal dopamine release | Demonstrated feasibility in operating room environment |
| Swine (Sus scrofa domesticus) | Wireless neurotransmitter monitoring [43] | Triangular waveform -0.4 to +1.5 V, 400 V/s, 10 Hz | Dopamine signaling responded sigmoidally to pulse intensity and pulse-width STN stimulation | Validated wireless systems for clinical application |
| Swine (Sus scrofa domesticus) | Adenosine during seizure termination [43] | Triangular waveform -0.4 to +1.5 V, 900 V/s, 10 Hz | Increased adenosine levels observed prior to seizure termination | Direct correlation with human patient recordings |
| Non-human Primate (Macaca sp.) | Reward-related dopamine signaling [43] | Various triangular waveforms -0.4/-0.6 V to +1.0/+1.4 V, 400 V/s, 10 Hz | Oxygen and pH changes associated with reward prediction; dopamine responses observed | Established behavioral correlates in complex learning tasks |
Protocol: FSCV in Swine Models for Adenosine Detection
Materials Required:
- Carbon fiber microelectrodes (7μm diameter, ~100μm exposed tip)
- Ag/AgCl reference electrode
- Stereotaxic surgical apparatus
- Voltammetric amplifier
- Data acquisition system
- Anesthesia delivery system
- Physiological monitoring equipment
Experimental Procedure:
Animal Preparation: Anesthetize swine using approved protocols (e.g., ketamine/xylazine induction followed by isoflurane maintenance). Secure animal in stereotaxic frame and maintain physiological monitoring throughout procedure.
Surgical Approach: Perform craniotomy at target coordinates relative to bregma. Maintain sterile conditions throughout the procedure.
Electrode Placement: Position carbon fiber working electrode in target region (e.g., striatum or cortex) and implant Ag/AgCl reference electrode in contralateral hemisphere.
FSCV Parameter Setup: Apply triangular waveform (-0.4 V to +1.5 V, 400 V/s) at 10 Hz frequency. Allow electrode to stabilize for 30-60 minutes before recording.
Signal Validation: Implement the "Five Golden Rules" for adenosine identification:
- Record background-subtracted cyclic voltammograms for adenosine signature confirmation.
- Perform electrical stimulation (300 μA, 60 Hz, 2 ms pulse width) to evoke adenosine release when applicable.
- Administer receptor-specific pharmacological agents (e.g., adenosine A1 receptor antagonist DPCPX) to verify signal identity.
Data Collection: Record spontaneous and evoked adenosine transients. Include appropriate controls for physiological confounds (pH changes, oxygen fluctuations).
Histological Verification: Upon experiment completion, perfuse animal and perform brain sectioning to verify electrode placement sites.
This protocol has been successfully implemented in multiple swine studies investigating adenosine dynamics, particularly in the context of deep brain stimulation and seizure activity [43]. The environment can be deliberately adapted to mimic human operating room conditions to enhance translational validity.
Human Feasibility Studies: Methodological Considerations
Technical Challenges in Human Adenosine Measurement
The transition from animal models to human studies introduces significant methodological challenges for adenosine detection. In blood measurements, the extremely short half-life of adenosine (<1 second) necessitates specialized sampling techniques to accurately determine circulating concentrations [59]. For CNS measurements using FSCV, additional considerations include electrode biocompatibility, safety monitoring during intraoperative recordings, and management of motion artifacts in conscious patients.
Recent research has highlighted that "advancing the knowledge of underlying mechanisms and biological processes associated with the complex chemistry of the human brain is critical to the development of new and improved therapeutic interventions" [43]. For example, clinical use of FSCV may allow characterization of neurochemical signaling evoked by deep brain stimulation (DBS), potentially advancing treatment of neurologic diseases. However, these potential benefits must be balanced against rigorous safety protocols and methodological validation.
Protocol: Accurate Measurement of Endogenous Adenosine in Human Blood
Materials Required:
- STOP solution (see Research Reagent Solutions below)
- S-Monovette K3E tubes (Sarstedt)
- Pre-chilled centrifuge
- UPLC-tandem mass spectrometry system
- Stable isotope labeled adenosine and AMP (Cambridge Isotope Laboratories)
- Inhibitors: AOPCP, EDTA, dipyridamole, NBMPR, EHNA, 5-Iodotubericidin
Experimental Procedure:
STOP Solution Preparation: Prepare STOP solution containing:
- AOPCP (200 μM final concentration) - inhibits CD73-mediated adenosine formation
- EDTA (4 mM) - chelator that inhibits nucleotidases
- Dipyridamole (10 μM) and NBMPR (10 μM) - inhibit ENT1-mediated adenosine uptake
- EHNA (10 μM) - inhibits adenosine deaminase
- 5-Iodotubericidin (50 nM) - inhibits adenosine kinase
- Prepare in PBS (pH 7.4) with stable isotope internal standards
Blood Collection Protocol:
- Add 1.3 mL STOP solution to 2.6 mL S-Monovette tubes
- Discard first 2 mL of venous blood to avoid contamination
- Collect blood directly into tube containing STOP solution
- Invert tube immediately 10 times for rapid mixing
Sample Processing:
- Centrifuge blood at 2000 × g for 10 minutes at 4°C within 5 minutes of collection
- Transfer plasma to pre-chilled tubes
- Add precipitation solvent (e.g., methanol) for protein removal
- Centrifuge and collect supernatant for analysis
UPLC-tandem MS Analysis:
- Use reverse-phase chromatography with HILIC or C18 columns
- Employ positive electrospray ionization with multiple reaction monitoring
- Quantify against calibration curves using stable isotope internal standards
- Maintain lower limit of quantification at 2 nmol/L
This optimized protocol enables accurate measurement of endogenous adenosine concentrations in human blood, with reported mean values of 13 ± 7 nmol/L in healthy volunteers [59]. The method has demonstrated utility in assessing pharmacological effects on adenosine concentration, such as the concentration-dependent ability of ticagrelor to conserve extracellular adenosine at clinically relevant exposures [59].
Protocol: Intraoperative FSCV in Human Patients
Materials Required:
- Clinical-grade carbon fiber microelectrodes
- Biocompatible reference electrode
- Voltammetric amplifier with clinical safety certification
- Data acquisition system integrated with surgical navigation
- Sterile electrode insertion apparatus
Experimental Procedure:
Ethical and Regulatory Compliance: Obtain IRB approval and informed consent. Clearly define research nature of procedure separate from therapeutic intervention.
Surgical Coordination: Coordinate with surgical team to integrate FSCV recordings with standard procedure (e.g., DBS electrode implantation).
Electrode Placement: Sterilely position working electrode using surgical guidance systems. Place reference electrode at extracranial site.
FSCV Parameter Application: Apply optimized waveform parameters (-0.4 V to +1.5 V, 400 V/s at 10 Hz) with appropriate electrical safety limits.
Signal Validation: Record background-subtracted cyclic voltammograms. Compare with library of known electrochemical signatures.
Physiological Monitoring: Continuously monitor patient physiological parameters throughout recording.
Data Interpretation: Apply conservative interpretation criteria, acknowledging potential confounds in clinical environment.
This protocol has been successfully implemented in limited human studies, such as those investigating adenosine release during seizure termination, where increased adenosine levels were observed just prior to seizure cessation [43] [55] [43].
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 3: Key Research Reagent Solutions for Adenosine Measurement
| Reagent/Category | Composition/Type | Function | Application Context |
|---|---|---|---|
| STOP Solution | AOPCP (200 μM), EDTA (4 mM), Dipyridamole (10 μM), NBMPR (10 μM), EHNA (10 μM), 5-Iodotubericidin (50 nM) | Prevents both formation and clearance of adenosine in blood samples | Human blood collection for accurate adenosine quantification [59] |
| Carbon Fiber Microelectrodes | Single carbon fiber (T-650, 7 μm diameter) sealed in glass capillary | Working electrode for FSCV measurements; enables detection of electroactive analytes | In vivo FSCV in animal models and human intraoperative recordings [11] |
| Adenosine Receptor Modulators | Selective agonists/antagonists for A1, A2A, A2B, and A3 receptors | Pharmacological validation of adenosine signals; investigation of receptor-specific effects | All experimental contexts for mechanistic studies |
| Stable Isotope Labels | 13C10-15N5-adenosine, 15N5-AMP | Internal standards for mass spectrometry quantification; tracking of adenosine dynamics | Analytical method validation; metabolic studies |
| Enzyme Inhibitors | EHNA (ADA inhibitor), 5-Iodotubericidin (AK inhibitor) | Inhibition of adenosine degradation pathways; increases extracellular adenosine half-life | Multiple experimental contexts to modulate adenosine signaling |
The translation of adenosine detection methodologies from basic research to clinical application requires a systematic, phased approach that leverages the unique strengths of large animal models and carefully designed human feasibility studies. The integrated pathway presented in this application note emphasizes validation at each stage, from initial technical development in porcine models to final implementation in clinical settings. As the field advances, continued refinement of these protocols will enhance the reliability and safety of adenosine measurements in human subjects, ultimately supporting the development of adenosine-targeted therapies for neurological and cardiovascular disorders.
The demonstrated ability of FSCV to detect rapid adenosine fluctuations in both animal models and human patients suggests that "adenosine transmission in the brain may have characteristics similar to those of classical neurotransmitters, such as dopamine and norepinephrine" [11]. This paradigm shift in understanding adenosine signaling underscores the importance of continued methodological development in this field, with large animal models serving as an indispensable component of the translational pathway.
Conclusion
Fast-Scan Cyclic Voltammetry has fundamentally advanced our understanding of rapid adenosine signaling in the brain, revealing its transient, activity-dependent release and regional neuromodulatory effects. Methodological innovations in waveforms, electrode design, and data analysis have continuously pushed the limits of sensitivity and selectivity. While challenges like biofouling and signal validation remain active areas of research, the integration of FSCV with other techniques and the development of more robust, biocompatible electrodes are paving a clear path forward. The future of FSCV for adenosine detection lies in its translation to clinical settings, where it holds immense promise for providing real-time neurochemical feedback in closed-loop neuromodulation systems for treating disorders like Parkinson's disease and epilepsy, ultimately enabling more personalized and effective therapeutic interventions.
References
- 1. Adenosine [https://en.wikipedia.org/wiki/Adenosine]
- 2. What is Adenosine? [https://www.news-medical.net/health/What-is-Adenosine.aspx]
- 3. Adenosine and Sleep: Understanding Your Sleep Drive [https://www.sleepfoundation.org/how-sleep-works/adenosine-and-sleep]
- 4. Fast-scan Cyclic Voltammetry for the Characterization of ... [https://www.sciencedirect.com/science/article/pii/S2001037014000567]
- 5. Adenosine, caffeine, and sleep–wake regulation: state of the ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9541543/]
- 6. Adenosine, energy metabolism and sleep homeostasis [https://www.sciencedirect.com/science/article/abs/pii/S1087079210000663]
- 7. Multiplexing FSCV and iGluSnFR3 Sensors Reveals ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12230777/]
- 8. Structural Similarity Image Analysis for Detection of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7478140/]
- 9. Improved longevity and in vivo performance of ... [https://www.frontiersin.org/journals/bioengineering-and-biotechnology/articles/10.3389/fbioe.2025.1579380/full]
- 10. Fast-scan Cyclic Voltammetry for the Characterization of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4720017/]
- 11. Real time adenosine fluctuations detected with fast-scan cyclic ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4651740/]
- 12. Improved longevity and in vivo performance of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12411426/]
- 13. Defining a Path Toward the Use of Fast-Scan Cyclic ... [https://www.frontiersin.org/journals/neuroscience/articles/10.3389/fnins.2021.728092/full]
- 14. Simultaneous Recognition and Detection of Adenosine ... [https://www.mdpi.com/1424-8220/24/20/6648]
- 15. Fast-Scan Cyclic Voltemmtery of Adenosine in Venton Lab [https://pineresearch.com/support-article/fast-scan-cyclic-voltemmtery-of-adenosine-in-venton-lab/]
- 16. Innovating carbon-based electrodes for direct neurochemical ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12364121/]
- 17. Carbon nanospikes have improved sensitivity and ... [https://uva.theopenscholar.com/venton-group/publications/carbon-nanospikes-have-improved-sensitivity-and-antifouling-properties-adenosine]
- 18. Carbon microelectrodes for the measurement of ... [https://www.frontiersin.org/journals/bioengineering-and-biotechnology/articles/10.3389/fbioe.2025.1569508/full]
- 19. An ultrasensitive electrochemical biosensor for ATP ... [https://www.sciencedirect.com/science/article/abs/pii/S0925400525008573]
- 20. Automated Algorithm for Detection of Transient Adenosine ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5312768/]
- 21. Sawhorse Waveform Voltammetry for Selective Detection ... [https://acs.figshare.com/articles/journal_contribution/Sawhorse_Waveform_Voltammetry_for_Selective_Detection_of_Adenosine_ATP_and_Hydrogen_Peroxide/2039469]
- 22. Comonitoring of adenosine and dopamine using the Wireless ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC2852872/]
- 23. Stabilized carbon coating on microelectrodes for scalable ... [https://www.nature.com/articles/s41467-025-58388-z]
- 24. Structural Similarity Image Analysis for Detection of Adenosine ... [https://figshare.com/articles/journal_contribution/Structural_Similarity_Image_Analysis_for_Detection_of_Adenosine_and_Dopamine_in_Fast-Scan_Cyclic_Voltammetry_Color_Plots/12682502?backTo=/collections/_/5068898]
- 25. Structural similarity index measure [https://en.wikipedia.org/wiki/Structural_similarity_index_measure]
- 26. Structural Similarity Index(SSIM) | by Abhishek Kumar ... [https://medium.com/@akp83540/structural-similarity-index-ssim-c5862bb2b520]
- 27. SSIM-Based Autoencoder Modeling to Defeat Adversarial ... [https://www.mdpi.com/1424-8220/24/19/6461]
- 28. SSIM: Structural Similarity Index [https://www.imatest.com/docs/ssim/]
- 29. AI-Assisted Detection of Biomarkers by Sensors and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11274923/]
- 30. Advancement of machine learning algorithms in biosensors [https://www.sciencedirect.com/science/article/abs/pii/S000989812500556X]
- 31. A Flexible Software Platform for Fast-Scan Cyclic Voltammetry ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3838858/]
- 32. Understanding the different effects of fouling mechanisms on ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11648937/]
- 33. Mitigating the Effects of Electrode Biofouling-Induced ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8281608/]
- 34. Recent Advances in Fast-Scan Cyclic Voltammetry - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC7028521/]
- 35. Electrochemical determination of adenosine by natural ... [https://www.sciencedirect.com/science/article/pii/S2214180418301211]
- 36. In Vitro Biofouling Performance of Boron-Doped Diamond ... [https://www.mdpi.com/2079-6374/13/6/576]
- 37. Adenosine detection in serum using a surface plasmon ... [https://pubs.rsc.org/doi/d4lp00059e]
- 38. A fast scan cyclic voltammetric digital circuit with precise ... [https://www.sciencedirect.com/science/article/abs/pii/S0003267021005705]
- 39. Ohmic Drop Part I: Effect on measurements ... [https://www.biologic.net/documents/ohmic-drop-effect-on-measurements-electrochemistry-battery-corrosion-application-note-27/]
- 40. Fast Scan Cyclic Voltammetry: Chemical Sensing in the Brain ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5750125/]
- 41. Cyclic Voltammetry - Data Analysis [https://www.basinc.com/manuals/EC_epsilon/Techniques/CycVolt/cv_analysis]
- 42. Progress Towards Biocompatible Intracortical Microelectrodes ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4428498/]
- 43. Defining a Path Toward the Use of Fast-Scan Cyclic ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8633532/]
- 44. A Review: Electrode and Packaging Materials for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7873964/]
- 45. Exploring Neurotransmitters through In Vivo vs. In Vitro ... [https://www.mdpi.com/1424-8220/24/2/647]
- 46. An Introduction to Electrochemical Methods in Neuroscience [https://www.ncbi.nlm.nih.gov/books/NBK1845/]
- 47. An Update of the Classical and Novel Methods Used for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4428024/]
- 48. Neurochemistry and electroanalytical probes [https://www.sciencedirect.com/science/article/abs/pii/S1367593102003745]
- 49. Continuous monitoring of adenosine and its metabolites ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6774626/]
- 50. A review of the effects of FSCV and microdialysis ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4437820/]
- 51. Intracerebral microdialysis of adenosine and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6220825/]
- 52. An improved LC–S/MS method for the quantitation of ... [https://www.sciencedirect.com/science/article/abs/pii/S0731708512002701]
- 53. Increased cortical extracellular adenosine correlates with ... [https://pubmed.ncbi.nlm.nih.gov/24483230/]
- 54. A high-performance fluorescent sensor spatiotemporally ... [https://www.nature.com/articles/s41467-025-59530-7]
- 55. Feasibility of large experimental animal models in testing ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8040081/]
- 56. “Lost in translation?” Animal research in the era of precision ... [https://translational-medicine.biomedcentral.com/articles/10.1186/s12967-025-06084-3]
- 57. Animal models for the study of adenosine receptor function [https://pubmed.ncbi.nlm.nih.gov/15389588/]
- 58. Human in vivo research on the vascular effects of adenosine [https://www.sciencedirect.com/science/article/abs/pii/S001429990800277X]
- 59. Accurate measurement of endogenous adenosine in human ... [https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0205707]